Introduction
Listeria monocytogenes, a gram-positive bacterium, is ubiquitously present in the environment [1, 2]. The bacterium can infect a wide range of cell types; cross the intestinal, placental, and blood-brain barriers; and cause listeriosis in humans [2, 3]. Listeriosis is associated with a large proportion of the public health and economic burden; almost 1500 listeriosis cases are reported annually in the United States, where listeriosis accounts for approximately 4% of hospitalisations and 19% of deaths were caused by L. monocytogenes infection [4, 5]. L. monocytogenes is a facultative intracellular bacterium, but can escape from autophagy. The ability of L. monocytogenes to establish an intracellular niche and evade immune surveillance is critical to the outcome of infection [6].
Macrophages are important innate immune cells that defend against invading pathogens [7]. A recent report showed that infection of macrophages with L. monocytogenes induces the expression of miR-21 in cells to regulate phagocytosis, thereby limiting the intracellular load of the pathogen [8]. miR-21a is a typically upregulated miRNA in many mammalian cell types after L. monocytogenes infection [9]. Additionally, miR-21a is reportedly upregulated in many cancers, and therefore is also known as an ‘oncomiR’ [10, 11]. More recent studies have confirmed that miR-21 is upregulated in many infectious or inflammatory diseases [12-15]. The induction of miR-21 has been proposed to represent a pivotal switch in the inflammatory response [16].
The miRNAs target RNA transcripts encoding functional proteins and play critical roles in regulating gene expression. Programmed cell death 4 (PDCD4, also known as MA-3) was first identified as a gene upregulated during apoptosis [17]. PDCD4 protein, characterised as a tumour suppressor protein [18], is targeted by miR-21a in various types of tumours [19, 20]. A recent study showed that the miR-21/PDCD4 axis plays a vital role in the process of efferocytosis carried out by macrophages [21]. In gastric epithelial cells, Helicobacter pylori can downregulate PDCD4 expression by inducing miR-21a [22]. L. monocytogenes is the causative agent for several systemic infections; however, whether L. monocytogenes infection in macrophages modulates the miR-21/PDCD4 axis remains to be determined.
In this study, we sought to elucidate the modulation of the miR-21/PDCD4 axis in L. monocytogenes infected macrophages. Our results showed that miR-21 expression is upregulated in macrophages during infection with L. monocytogenes. The upregulated miR-21 downregulated the expression of PDCD4 during the late stage of L. monocytogenes infection. Further, the downregulation of PDCD4 was accompanied by activation of c-Jun and STAT3.
Material and methods
Cell culture
The mouse macrophage cell line RAW264.7 was obtained from the laboratory of Professor Wan of the Army Medical University of China. Cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM) (Gibco, MA, USA) supplemented with 10% (v/v) FBS (Lonza, Basel, Switzerland) and with Pen/Strep (10,000 µg/ml) (Lonza, Basel, Switzerland) in a humified incubator (> 95% humidity) set at 37oC and 5% CO2.
Bacterial strains
The L. monocytogenes wild-type strain (10403S) was provided by Professor Wan of the Army Medical University of China. L. monocytogenes were cultured in BHI broth (Sigma, USA) in a shaking incubator (200 rpm) set at 37°C. Bacteria were subsequently washed with Hanks Balanced Salt Solution (HBSS) (Gibco, MA) at 3000 rpm for 10 min. The growth of L. monocytogenes was monitored by measuring absorbance at 600 nm using a spectrophotometer.
Listeria monocytogenes infection
Infection of macrophages with L. monocytogenes was performed, as previously described [23]. Briefly, RAW264.7 cells (1 × 105 cells per well) were seeded in 24-well plates in the antibiotic-free complete medium. Subsequently, the cells were infected with L. monocytogenes at the multiplicity of infection (MOI) of 20 or 100. After infection, the plates were centrifuged at 1500 rpm for 15 min. Forty-five minutes later, cells were treated with 100 µg/ml gentamicin for 30 min to kill extracellular bacteria. To prevent re-infection, the cells were treated with 5 µg/ml gentamicin for up to four hours. For colony-forming units (CFU) assay, cells were lysed with 0.1% Triton X-100 (Sigma, MO, USA) and serially diluted. The diluted lysed cultures were spread on a BHI agar plate and incubated overnight at 37°C. The CFU of L. monocytogenes were determined as per the formula: CFU = number of colonies × dilution.
Treatment with miR-21a mimic
Mimics for miR-21a (miR-21a-5p) and control mimics were purchased from RiboBio (RiboBio Co. Ltd., Guangzhou, China).
PDCD4 silencing
Three different siRNAs targeting mouse PDCD4 mRNA were used in this study, and the sequences are listed in Table 1. Control siRNAs were purchased from RiboBio (siN0000001-1-5, RiboBio Co. Ltd., Guangzhou, China).
Cell transfection
RAW264.7 cells (1 × 105 cells per well) were seeded in 24-well plates in an antibiotic-free complete culture medium. The miR-21a mimic or siRNAs and the control siRNAs were transfected into cells using Lipofectamine 3000 reagent (Thermo Fisher Scientific, MA, USA) according to the manufacturer’s instructions. Cells were cultured for 48 h after transfection and subsequently used for bacteria burden assay.
RNA isolation and real-time quantitative PCR
Total RNA was isolated using RNA MicroPrep (ZYMO research, CA) according to the manufacturer’s protocol. Reverse transcription was performed using the High Capacity cDNA Reverse Transcriptase kit (Applied Biosystems, CA, USA). Quantitative real-time polymerase chain reaction (qRT-PCR) was performed in a CFX96 Real-Time PCR system (Bio-Rad, USA) using SYBR green qPCR Mix (TaKaRa, Japan). The expression level of miR-21a was normalised with U6 small nuclear RNA. Primer sequences used for these experiments are listed in Table 2.
Western blot analysis
Cells were lysed using RIPA lysis buffer (Thermo Fisher Scientific, MA, USA) containing Protease Inhibitor Cocktail (Cell Signalling Tech, MA, USA). The total protein content was quantified using the BCA protein assay kit (Thermo Fisher Scientific, MA, USA). Subsequently, proteins were separated using polyacrylamide gel electrophoresis and transferred onto polyvinylidene fluoride (PVDF) membranes (Millipore Corp, MA, USA). The membranes were blocked with 5% w/v bovine serum albumin in TBST buffer (Sigma, USA) for 1 h at room temperature and incubated with anti-PDCD4 and anti-β-actin antibodies (1 : 1000, Proteintech, USA) overnight at 4°C. The next day, the membranes were incubated with HRP linked goat anti-rabbit IgG (1 : 5000, Sigma, USA) for 40 min at room temperature and then washed three times with TBS-Tween20. The protein bands were visualised using Pierce TMECL Western Blotting Substrate (Thermo Fisher, MA, USA) and the digital images were captured using the ChemiDoc Touch Imaging System (Bio-Rad).
Phagocytic activity assay
The phagocytic activity of RAW264.7 cells was assessed using the FluoSpheres size kit #2 (Thermo Fisher, F8888, USA) according to the manufacturer’s instructions. Briefly, dendritic cells (DCs) were plated (1 × 105 cells/well) into 6-well plates and cultured for 12 h. Next, DCs were infected with FluoSpheres beads at the MOI = 20 for 1 h. The DCs were finally washed with phosphate-buffered saline (PBS) and subjected to flow cytometric analysis.
Results
PDCD4 was down-regulated in macrophages at the late stage of Listeria infection
An MOI of 100 : 1 is commonly used for bacterial burden assay using L. monocytogenes. Our results showed that the macrophage had a high bacterial burden 2 h after L. monocytogenes infection. After infection for 5 h there was a marginal increase (13.66%) in the L. monocytogenes burden in macrophages (Fig. 1A). Further, we observed that the expression of miR-21a increased consistently until 5 h after L. monocytogenes infection in macrophages (Fig. 1B). These results were consistent with a previously reported finding that the expression of miR-21a was significantly upregulated 2 h post-infection [8].
Fig. 1
PDCD4 is downregulated at the late stage of Listeria infection in macrophages. A) RAW264.7 cells were infected with L. monocytogenes at MOI = 100. The number of intracellular bacteria was determined at 2 and 5 h post-infection and expressed as log CFU/ml. B) At 5 h post-infection, miR-21a expression was assayed by qPCR in RAW264.7 cells. C, D) RAW264.7 cells were infected with L. monocytogenes at MOIs of 20 and 100. PDCD4 protein expression was examined using western blot at 2 h (C) or 5 h (D) post-infection (**p < 0.01)
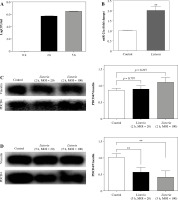
As a downstream target of miR-21a, the expression of PDCD4 has been implicated in the induction of anti-inflammatory phenotype in macrophages [21]. To assess the role of PDCD4 during macrophage infection of L. monocytogenes, we examined the expression of PDCD4 protein. Our results showed that the expression of PDCD4 protein decreased significantly 5 h after L. monocytogenes infection in macrophages. However, interestingly, we observed that the expression of PDCD4 protein did not decrease significantly 2 h after infection (Fig. 1C, D). Similar results were obtained when we used MOI of 20 : 1 for macrophage infection (Fig. 1C, D). Overall, these results indicated that the miR-21a/PDCD4 axis or PDCD4 protein might play a role only during the late stage of L. monocytogenes infection in macrophages.
Knockdown of PDCD4 contributed to enhanced Listeria infection in macrophages
To further validate the involvement of the miR-21a/PDCD4 axis in determining L. monocytogenes burden, we transfected miR-21a mimic to overexpress miR-21a in macrophages, while scrambled oligonucleotides were used as a control. Twenty-four hours later, the transfected macrophages were infected by L. monocytogenes at the MOI of 100 : 1. After 5 h, CFU assays were performed to evaluate the L. monocytogenes burden in macrophages. Our results showed a significant increase in L. monocytogenes burden in macrophages with overexpression of miR-21a (Fig. 2A). Next, we depleted PDCD4 by transfecting macrophages with siPDCD4; siNC (siN0000001-1-5, RiboBio) was used as a control. The transfection eficiency was determined using western blot. The results showed that PDCD4 protein expression was remarkably downregulated in macrophages after 48 h of transfection with siPDCD4 (Fig. 2C). As expected, macrophages with downregulated PDCD4 presented a higher L. monocytogenes burden (Fig. 2B). These results suggested that the upregulation of miR-21a in macrophages could enhance phagocytosis of L. monocytogenes and depended on PDCD4 expression.
Fig. 2
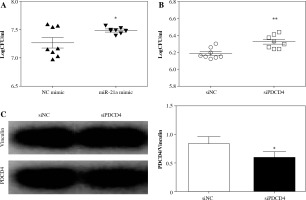
PDCD4 silencing contributed to a higher L. monocytogenes burden in macrophages. RAW264.7 cells were transfected with 20 nM miR-21a mimic or siRNA of PDCD4. Over 48 h, cells were infected with L. monocytogenes at MOI of 100. A, B) The number of intracellular bacteria was determined at 5 h post-infection and expressed as logCFU/ml. C) At 5 h post-infection, PDCD4 protein was examined using western blot (*p < 0.05, **p < 0.01)
PDCD4 did not participate in the Listeria uptake by macrophages
We next determined the mechanism behind the high L. monocytogenes burden in macrophages. RAW264.7 cells were transfected with siPDCD4 or control siNC. After 48 h, transfected cells were incubated with FluoSpheres carboxylate-modified microspheres (yellow-green fluorescent) for 5 h. Finally, the cells with microspheres were quantified using FACS. As shown in Figure 3A and B, there was no significant difference in the percentage of microsphere-positive cells whether PDCD4 was depleted or not. Furthermore, we also examined and calculated the mean fluorescence intensity of the RAW264.7 cells and found no significant difference between PDCD4 depleted or WT RAW264.7 cells (Fig. 3A and C). The transfection eficiency in these experiments was determined using western blots (Fig. 3D), and the results indicated that the depletion of PDCD4 did not affect the uptake of L. monocytogenes by macrophages. In other words, PDCD4 might be involved in the engulfment, rather than uptake, of L. monocytogenes.
Fig. 3
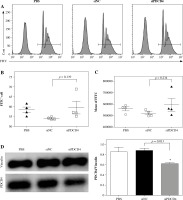
PDCD4 silencing did not change the uptake ability of RAW264.7 cells. RAW264.7 cells were transfected with 20 nM siPDCD4 or control siNC. Over 48 hours, cells were incubated with FluoSpheres carboxylate-modified microspheres for another 5 h. A) The cells with microspheres were quantified using FACS. The percentage (B) and the mean fluorescence intensity (C) of microsphere-positive cells were calculated. D) At 5 h post-infection, PDCD4 protein was examined using western blot (*p < 0.05)
Phosphorylation of c-Jun and STAT3 might enhance intracellular bacteria burden
A previous report showed that miR-21a was involved in the successful engulfment of apoptotic cells by wound-associated macrophages via the PDCD4/c-Jun pathway [21]. To determine the mechanism of miR-21a/PDCD4-mediated engulfment of L. monocytogenes, we examined the phosphorylation of c-Jun in L. monocytogenes infected RAW264.7 cells. As shown in Figure 4A, the phosphorylation of c-Jun was significantly increased in RAW264.7 cells infected with L. monocytogenes at the MOI of 20 : 1 and 100 : 1. These results indicated that the miR-21a/PDCD4 axis triggered c-Jun activation in L. monocytogenes infected RAW264.7 cells.
Fig. 4
Phosphorylated c-Jun and STAT3 were elevated during the late stage of Listeria infection in macrophages. A) RAW264.7 cells were infected with L. monocytogenes at MOI of 100. At 5 h post-infection, protein expression of PDCD4, STAT3, phosphorylated STAT3, c-Jun and phosphorylated c-Jun was examined using western blot. B) Schematic representation of the mechanism by which PDCD4 could modulate L. monocytogenes infection in macrophages. At the late stage of infection, overexpression of miR-21a leads to PDCD4 downregulation which results in the activation and phosphorylation of c-Jun/STAT3. Activated c-Jun/STAT3 is likely to upregulate the transcription of inflammatory genes such as AP-1, IL-6, IL-1β and downregulate IL-10 expression, thereby possibly contributing to modulation in macrophage-mediated immune response (*p < 0.05, **p < 0.01)
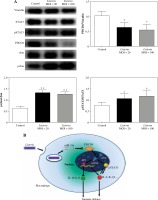
As the interaction between STAT3 and c-Jun mediates the downstream signalling [24, 25], we further examined the expression of phosphorylated STAT3 in L. monocytogenes infected RAW264.7 cells. Consistent with the phosphorylation of c-Jun, the phosphorylation of STAT3 was significantly increased as well in RAW264.7 cells infected with L. monocytogenes (Fig. 4A). Collectively, these results suggested that the activation of c-Jun and STAT3 regulates miR-21a/PDCD4 axis, thus impacting L. monocytogenes engulfment by macrophages.
Discussion
Macrophages are the effector cells of the innate immune system and are involved in defence against invading pathogens, and secrete inflammatory mediators [7]. Enhanced expression of miR-21a was reportedly associated with the immune response against macrophage infections caused by L. monocytogenes [8], Mycobacterium tuberculosis [26], and LPS treatment [27] of macrophages. In this study, we found that infection with L. monocytogenes upregulated the expression of miR-21a in macrophages. We observed that the high expression of miR-21a was maintained even at the late stage of L. monocytogenes infection (Fig. 1B). This observation was consistent with a previously published finding that the expression of miR-21a was significantly increased at 2 h post-infection [8]. As miR-21 has been implicated in the macrophage polarisation and inflammatory response [28], our results suggest that miR-21a might play an important role during the late stage of L. monocytogenes infection in macrophages.
PDCD4 has been characterised as a tumour suppressor protein [18] and is targeted by miR-21a in many types of tumours [19, 20]. Interestingly, in our study, we found that the expression of PDCD4 protein in macrophages decreased 5 h, but not 2 h, after L. monocytogenes infection (Fig. 1C, D). Further, our results showed that the macrophages with silenced PDCD4 presented a higher L. monocytogenes burden (Fig. 2B), thereby validating our previous findings. Together, these results indicated that PDCD4 was involved in L. monocytogenes pathogenesis in macrophages. The bacterium invades macrophages to establish an intracellular niche and evade the host immune response. The pathogen has evolved to escape from the phagolysosome and instead grows and replicates within the macrophage cytoplasm [6]. The results from the FluoSpheres carboxylate-modified microspheres incubation showed that PDCD4 did not affect macrophages’ ability to take up L. monocytogenes (Fig. 3). So far, there have been no reports indicating the involvement of PDCD4 in the macrophage-mediated endocytosis of bacteria. However, our results show that the PDCD4 might be involved in the engulfment of L. monocytogenes by macrophages. Previously, the PDCD4/c-Jun pathway has been implicated in the successful engulfment of apoptotic cells by wound-associated macrophages [21]. Transcription factors STAT3 and c-Jun have been shown to modulate many important cellular processes, through the regulation of gene expression in tumour cells [24, 25]. Here, we analysed the potential pro-apoptotic effector role of PDCD4 in macrophages infected by L. monocytogenes. Our results showed that the downregulation of PDCD4 protein was accompanied by the phosphorylation and activation of both c-Jun and STAT3 during the late stage of L. monocytogenes infection in macrophages (Fig. 4A). In a previous study, PDCD4-mediated inhibition of activator protein-1 (AP-1) showed concomitant suppression of c-Jun phosphorylation [29, 30]. This is significant since c-Jun activation plays an important role in the inflammatory signalling of macrophage function [31]. While PDCD4 is pro-apoptotic, STAT3 has anti-apoptotic [18, 32] functions. The downregulation of PDCD4 triggers STAT3 activation, which activates the inflammatory cascade, leading to IL-6 expression in L. monocytogenes infected macrophages.


