Introduction
Long non-coding RNA (lncRNA) was first identified by Okazaki et al. [1] in 2002 when they conducted large-scale sequencing of mouse cDNA. lncRNAs are non-protein coding transcripts (ncRNA) longer than 200 nucleotides. Recent studies have revealed that lncRNA play important roles in cell development and metabolism [2], especially in miRNA regulation and protein encoding. Furthermore, a variety of cancers have abnormal expression of specific lncRNA, which can form ribonucleoprotein complexes and interact with proteins to alter the biobehavior of cancer cells [3]. Their putative role in the occurrence and development of cancers has also been suggested [4].
Analysis of the lncRNA expression profile in normal oral mucosa and oral precancerous lesions revealed 325 differentially expressed lncRNA, 60% of which had abnormal expression in the oral precancerous lesions [5]. Metastasis-associated lung adenocarcinoma transcript 1 (MALAT-1) was the first lncRNA found to be related to cancer metastasis [5]. Ji et al. [6] found that MALAT-1 was highly expressed in non-small cell lung cancer (NSCLC) with early metastasis, and high MALAT-1 expression predicted a poor prognosis of NSCLC. In liver cancer, MALAT-1 played an important regulatory role in its occurrence and development [7], and it is upregulated in hepatocellular carcinoma (HCC), colon cancer, pancreatic cancer, breast cancer and prostate cancer [8–10]. Increased MALAT-1 expression in pharyngeal squamous cell carcinoma (SCC) was also demonstrated, increasing dramatically in stage T3 or T4 disease compared with stage T1 or T2 [11].
In addition to MALAT-1, roles for other lncRNA in cancer progression have been reported. Specifically, high UCA1 expression in bladder cancer cells increased their proliferation and resistance to therapies [12]. In breast cancer, UCA1 inhibited the tumor suppressor gene P27 [13]; UCA1 expression was also correlated with lymph node metastasis in patients with tongue SCC (TSCC) [14]. In addition, another lncRNA, BC200, was overexpressed in lung cancer, breast cancer, cervical cancer and ovarian cancer [15]. Furthermore, steroid receptor RNA activator (SRA) may modulate androgen receptor and estrogen receptor activity; therefore, it may play an important role in breast cancer [16].
Because few studies have evaluated the expression of lncRNA in oral SCC (OSCC) [17], this study aimed to investigate the expression of MALAT-1, BC200, UCA1 and SRA in OSCC and examine their possible role in the occurrence and development of OSCC. These particular lncRNAs were selected because each has been shown to be differentially expressed in a wide variety of cancers, including oral cancers [18]. Specifically, MALAT-1 was overexpressed in OSCC, and its suppression in OSCC cell lines reduced epithelial mesenchymal transition, migration, and invasion as well as tumor growth in vivo [19]. Similarly, UCA1 levels were upregulated in tongue squamous cell carcinoma, and its silencing reduced OSCC proliferation and metastasis by altering WNT/β-catenin signaling [20]. Although BC200 and SRA have not been reported in head and neck cancer, previous studies have shown that BC200 lncRNA is a potential predictive marker of poor prognosis in esophageal squamous cell carcinoma patients [21], and SRA is upregulated following vitamin D receptor ablation in skin cancer [22]. In addition, lncRNA SRA variants are associated with breast cancer risk [23]. We first examined the expression profile of MALAT-1, BC200, UCA1 and SRA in oral precancerous lesions by quantitative RT-PCR. Furthermore, the impact of MALAT-1 silencing on OSCC proliferation, apoptosis, colony formation capability and migration was examined in SCC4 cells.
MATERIAL AND METHODS
Collection of OSCC and adjacent normal tissues
In total, 14 pairs of OSCC tissues and corresponding adjacent normal tissues were collected from inpatients who received surgical interventions in the Department of Stomatology at the Changhai Hospital. None of the patients received chemotherapy and radiotherapy before the surgery, and patients with recurrent OSCC were excluded. The age, gender, location of primary cancer, clinical stage and neck lymph node metastasis were recorded. This study was approved by the Ethics Committee of the Changhai Hospital, and informed consent was obtained from each patient before enrollment in the study.
The cancer tissues and adjacent normal tissues were collected during the surgery, and a fraction of each was processed for pathological examination to identify the cancer tissues and normal tissues under a light microscope. The remaining tissues were stored at –80°C for further analysis.
Quantitative RT-PCR
Total RNA was isolated from frozen OSCC and adjacent normal tissues (about 100 mg) by the addition of 1 ml of Trizol reagent (Invitrogen, Carlsbad, CA, USA). In addition to 1000 ng of RNA, the reverse transcription reaction (20 μl total) included the following: 4 μl of 5 × PrimeScript Buffer, 1 μl of PrimeScript RT Enzyme Mix, 1 μl of 50 μM OligodT Primer, 1 μl of 100 μM random hexamers, and RNase-free deionized water (all from TaKaRa, Japan). The conditions for reverse transcription included 37°C for 15 min, 85°C for 5 s, and 4°C. Detection of MALAT-1, BC200, UCA1 and SRA expression was undertaken with an ABI7500 real-time PCR instrument (ABI, Foster City, CA, USA) and the primers shown in Table I (synthesized by Shanghai Sangon Biotech Co., Ltd, China). BRYT Green dye (Promega, Madison, WI USA) was used for the RT-PCR analysis, and the 20-μl reaction included the following: 10 μl of Go Taq qPCR Master Mix, 0.4 μl of forward primer (10 μM) and 0.4 μl of reverse primer (10 μM), 0.2 μl of 100× CXR, 2 μl of cDNA and 7 μl of nuclease-free water. Detection was performed in triplicate, and β-actin served as an internal control. The conditions for PCR were as follows: predenaturation at 95°C for 2 min, and then 40 cycles of 95°C for 15 s and 60°C for 1 min. The relative expression of each gene was calculated using the 2–ΔΔCt method.
Table I
Primers used for the quantitative RT-PCR analysis
Cell culture and siMALAT-1 transfection
We purchased SCC4 cells, a human tongue squamous cell carcinoma cell line derived from a 55-year-old man, from the ATCC (ATCC No. CRL-1624; Manassas, VA, USA). SCC4 cells were maintained in α-MEM containing 10% fetal bovine serum (FBS; both from GIBCO, Shanghai) at 37°C in an environment containing 5% CO2 at 95% humidity.
Lipofectamine 2000 (Invitrogen) was used for the transfection of siMALAT-1 to inhibit MALAT-1 expression. Four pairs of MALAT-1-siRNA and a scrambled control were synthesized in Genepharma Biotech Co., Ltd. (China): MALAT-1-siRNA-3097, 5′-GAGGUGUAAAGGGAUUUAUTT-3′ and 5′-AUAAAUCCCUUUACACCUCTT-3′; MALAT-1-siRNA-5639, 5′-GGCAUUUGCAUCUUUAAAUTT-3′ and 5′-AUUUAAAGAUGCAAAUGCCTT-3′; MALAT-1-siRNA-6108, 5′-CCCUCUAAAUAAGGAAUAATT-3′ and 5′-UUAUUCCUUAUUUAGAGGGTT-3′; MALAT-1-siRNA-6960, 5′-GGCAAGUGGAAAUGUUUAATT-3′ and 5′-UUAAACAUUUCCACUUGCCTT-3′; and Scrambled siRNA, 5′-UUCUCCGAACGUGUCACGUTT-3′ and 5′-ACGUGACACGUUCGGAGAATT-3′.
After transfection of SCC4 cells with each siMALAT-1 pair as well as the scrambled control, MALAT-1 expression was detected by RT-PCR and compared with untransfected cells at 24 and 48 h after transfection. Briefly, the reverse transcription reaction included 2 μl of RNA, 10 μl of 2× reverse transcription buffer, 0.4 μl of MMLV reverse transcriptase (200 U/μl), 5 μl of random primer N6 (100 μM), 3 μl of dNTPs (2.5 mM), and 0.15 μl of RNasin (40 U/μl) (all from XX). The conditions for reverse transcription included 37°C for 60 min, 85°C for 10 min, and 4°C. The 20-μl reaction for the quantitative RT-PCR included the following: 10 μl of 2× quantitative PCR Master Mix, 0.08 μl of forward primer (20 μM) and 0.08 μl of reverse primer (20 μM), 2 μl of cDNA, 0.4 μl of Taq DNA polymerase (2.5 U/μl) and nuclease-free water to 20 μl. The conditions for PCR were as follows: predenaturation at 95°C for 3 min, and then 40 cycles of 95°C for 15 s and 62°C for 30 s and 72°C for 30 s.
Detection of SCC4 cell proliferation
SCC4 cell proliferation was detected with a CCK-8 kit (Dojindo, Japan) following the manufacturer’s instructions. Briefly, the cells were seeded into 96-well plates at a density of 2 × 104 cells/well and transfected the following day as described above. At 24 h after transfection with scrambled siRNA and siMALAT-1 6108 and 6960, cells were re-suspended in serum-free medium (GIBCO) and then incubated at 37°C in an environment with 5% CO2 for 24 h. At 24, 48, and 72 h, 10 μl of CCK-8 solution was added to each well, and cells were incubated for 1 h. Absorbance was measured in a microplate reader at 450 nm.
Detection of SCC4 cell apoptosis
Annexin V/PI double staining was performed to detect the apoptotic cells. After cell density was adjusted to 1 × 106/ml, 100 μl of the cell suspension was mixed with 5 μl of Annexin V-FITC (mbchem). After the cells were incubated at room temperature in the dark for 5–15 min, 5 ml of propidium iodide (PI) was added, and the cells were incubated in the dark at room temperature for 5–15 min. Following addition of 400 μl of buffer, the cell suspension was subjected to flow cytometry at 480 nm.
Measurement of anchorage-independent growth of SCC4 cells
For the lower gel, 1.2% agarose was mixed with α-MEM containing 20% FBS at a ratio of 1 : 1, and 1.5 ml of this mixture was added to each well of a 6-well plate, which was then allowed to stay at room temperature until the agarose solidified. For the upper gel, 0.7% agarose was mixed with α-MEM containing 20% FBS at a ratio of 1 : 1, followed by addition of approximately 1000 transfected cells. This mixture was added to the pre-coated dishes. When the upper gel solidified, the dishes were incubated at 37°C in an environment with 5% CO2. At 5, 6 and 7 days, three fields were randomly selected, the colonies were counted, and the average was calculated.
Detection of SCC4 cell migration
Cell migration was determined using Transwell chambers (pore size: 8 μm; Corning, Ithica, NY, USA). A cell suspension of approximately 1 × 105 cells in 100 μl was added to the upper chamber of a Transwell chamber. Then, 500 μl of α-MEM containing 20% FBS was added to the lower chamber. After 48 h, a swab was used to collect the cells in the upper chamber, and 4% paraformaldehyde was added to the lower chamber to fix the cells in the chamber for 20 min. After drying, crystal violet was added to the lower surface of the filter of the Transwell chamber, followed by incubation at room temperature for 30 min. After washing in PBS once, cells were observed under an inverted microscope and photographed.
Statistical analysis
Data were presented as mean ± standard deviation for normally distributed continuous variables or median (inter-quartile range, the range between the 25th percentile and the 75th percentile) for non-normally distributed scale parameters. The number and percentage were calculated for categorical variables. To examine the difference between groups, paired t-tests were carried out for comparisons between normal and cancer tissues; Wilcoxon signed rank tests were used when continuous variables had skewed distributions. A general linear model, including time, group, and interaction between these two parameters, was built to test whether the group differences were altered by time points or vice versa. Once a significant interaction between time and group was identified, the group difference at each time point and time effect within each group were separately examined by one-way analysis of variance (ANOVA). ANOVA was also used to compare between four or more groups. Bonferroni’s method for variables with equal variances across groups or Dunnett’s T3 test for parameters with unequal variances was then implemented after the ANOVA revealed significant findings. P < 0.05 was considered significant. All statistics were two-sided, and the analyses were performed by SPSS software (version 15.0, SPSS Inc., Chicago, IL, USA).
RESULTS
OSCC patient characteristics
A total of 14 patients were enrolled in this study. As shown in Table II, the average age of the study population was 55.1 ±17.0 years (range: 25–85 years). Most of the patients were male (64.3%) with stage I OSCC (57.1%). In addition, most of the samples were highly differentiated (71.4%), and half of the patients had OSCC of the tongue (Table II).
Expression of MALAT-1, BC200, UCA1 and SRA in OSCC and normal oral mucosa
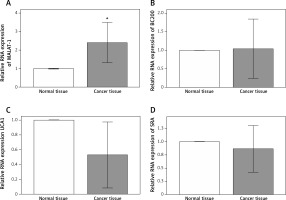
Because previous studies have reported putative roles for MALAT-1, BC200, UCA1 and SRA in other cancer types [5–17], we first analyzed their expression in 14 OSCC tissues and compared it with adjacent normal tissues. Compared with normal tissue, OSCC tissue showed a significantly increased MALAT-1 RNA expression (p = 0.004; Figure 1). However, no differences in BC200, UCA1 and SRA RNA expression were observed (Figure 1). The differential expression of MALAT-1 between the OSCC and adjacent normal tissues suggests that it may have a role in OSCC progression. Thus, further studies focused on MALAT-1.
The effects of MALAT-1 suppression on SCC4 cell proliferation and apoptosis
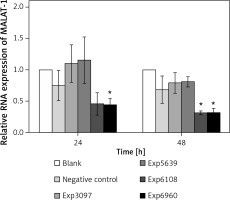
To determine the impact of MALAT-1 in OSCC, we first suppressed its expression in SCC4 cells by siRNA. The inhibitory effects of four siRNAs (number 3097, 5639, 6108, and 6960) were first compared. As shown in Figure 2, only siRNA 6960 significantly suppressed MALAT-1 RNA expression as compared to the untransfected, blank group (p = 0.009). After 48 h, both siRNA 6108 and 6960 significantly inhibited MALAT-1 gene expression as compared to the blank group (both p ≤ 0.016; Figure 2). Therefore, subsequent experiments only used 6108 and 6960 siRNA.
Figure 2
siRNA-mediated suppression of the MALAT-1 gene. The efficiency of MALAT-1 silencing was analyzed at 24 and 48 h following transfection in SCC4 cells by quantitative RT-PCR
Data were presented as mean ± standard deviation and tested by analysis of variance (n = 3 per group). *p < 0.05, significantly different from the blank group.
We next determined the impact of MALAT-1 knockdown on SCC4 proliferation and apoptosis. Although no differences were seen at 24 and 48 h, siRNA 6108 significantly inhibited SCC4 proliferation at 72 h as compared to the blank group (p = 0.004; Figure 3). In addition, silencing MALAT-1 expression with either 6108 or 6960 siRNAs significantly increased the apoptosis rates of SCC4 cells as compared to the blank group (both p < 0.001; Figures 4 A and B). Thus, MALAT-1 expression alters SSC4 proliferation and apoptosis.
Figure 3
Effects of MALAT-1 knockdown on proliferation of SCC4 cells (n = 5 per group). Data were presented as mean ± standard deviation. Comparisons between groups at each time point or between time points within a given group are all tested by analysis of variance
*p < 0.05, significantly different from the blank group.
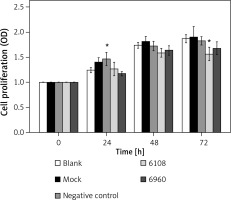
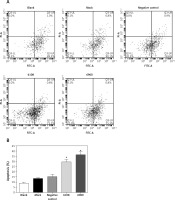
Figure 4
Effects of MALAT-1 knockdown on SCC4 cell apoptosis. A – Representative flow cytometry data from each group are shown. B – Apoptosis of SCC4 cells (n = 3 per group). Data were presented as mean ± standard deviation. Comparisons between groups at each time point or between time points within a given group are all tested by analysis of variance
*p < 0.05, significantly different from the blank group.
Inhibition of MALAT-1 expression reduces SCC4 cell anchorage-independent growth and migration
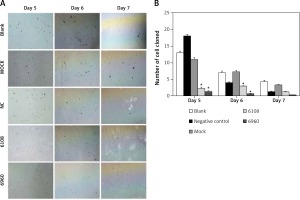
Previous studies on other cancer types have indicated a role for MALAT-1 in metastasis [5, 6]. Therefore, we first examined the impact of MALAT-1 suppression on anchorage-independent growth using a soft agar colony formation assay (Figure 5 A). As shown in Figure 5 B, the number of clones was significantly reduced following silencing with 6108 or 6960 as compared to the blank group at days 5 and 6, but not at day 7.
Figure 5
Effects of MALAT-1 knockdown on anchorage- independent growth of SCC4 cells. A – Representative SCC4 cell colony growth in each group at days 5–7. B – Anchorage-independent growth of SCC4 cells (n = 3 per group). Data were presented as mean ± standard deviation. Comparisons between groups at each time point or between time points within a given group are all tested by analysis of variance. *p < 0.05, significantly different from the blank group
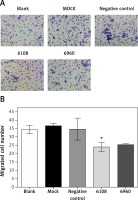
We next used a Transwell chamber to evaluate the effect of siMALAT-1 on SCC4 cell migration (Figure 6 A). At 48 h following MALAT-1 silencing, siRNA 6108 significantly reduced cell migration as compared to the blank group (p = 0.004; Figure 6 B). Although cells transfected with siRNA 6960 also had a lower amount of migrating cells than the blank group, the difference did not reach statistical significance. Taken together, these data suggest that MALAT-1 may also promote OSCC metastasis through enabling anchorage-independent growth and promoting migration.
Figure 6
Effects of MALAT-1 knockdown on migration of SCC4 cells. A – Representative SCC4 cell migration in each group. B – Migration of SCC4 cells using Transwell culture systems (n = 3 per group). Data were presented as mean ± standard deviation. Comparisons between groups at each time point or between time points within a given group are all tested by analysis of variance. *P < 0.05, significantly different from the blank group
Discussion
Although previous studies have shown a role for lncRNA in cancer progression and metastasis [24], few studies have assessed their expression and function in OSCC [17]. In the present study, MALAT-1 was differentially expressed between the OSCC and adjacent normal tissue, which is consistent with Fang et al. [14], who analyzed 94 TSCC samples and observed increased MALAT-1 expression. Furthermore, its suppression via siRNA resulted in increased SCC4 cell apoptosis as well as reduced proliferation, anchorage-independent growth, and migration. These studies suggest that increased MALAT-1 expression in OSCC may promote tumor cell growth and metastasis.
Analysis of two TCGA databases for head and neck cancer that included the four lncRNAs we analyzed in the present study (MALAT-1, UCA1, BC200 and SRA) revealed that MALAT-1 had a higher degree of gene amplification in 530 samples as compared with the other three lncRNAs. Consistent with our data showing a reduction in anchorage-independent growth and migration of SCC4 cells following MALAT-1 suppression, previous studies have suggested an association of MALAT-1 with metastasis and ultimately patient survival. Specifically, in patients with pharyngeal squamous cell carcinoma, increased MALAT-1 expression was observed in those with neck lymph node metastasis [11]. High MALAT-1 expression was also detected in nasopharyngeal carcinoma 5–8F cells with high metastatic potential [25].
Several studies have examined the mechanism by which MALAT-1 may promote tumor metastasis. In a manner similar to that described for the lncRNA HOTAIR, which decreases E-cadherin expression to promote OSCC metastasis [26], MALAT-1 overexpression in nasopharyngeal carcinoma CNE-1 cells by RNA activation inhibited epithelial cadherin and vimentin expression, which was accompanied by elevated invasion and metastasis [25]. In bladder cancer cells, MALAT-1 downregulation suppressed the expression of genes involved in epithelial-mesenchymal transition (EMT) [27]. In addition, the regulation of genes related to cell motility, including CTHRC1, CCT4, HMMR and ROD1, by MALAT-1 has been shown [28], which is similar to recent studies in which lncRNAs regulated matrix metalloproteinase expression in OSCC [29, 30]. Thus, further studies will examine the mechanism through which MALAT-1 promotes SSC4 cell migration.
In addition to impacting cell migration, MALAT-1 silencing reduced SSC4 cell proliferation and increased their apoptosis. This is consistent with data from CNE-1 cells in which MALAT-1 overexpression increased their proliferation [25]. However, the mechanism by which MALAT-1 influences cell growth and apoptosis remains unknown.
Once MALAT-1 is synthesized by RNA polymerase II, the products localize to nuclear speckles. Intranuclear MALAT-1 produces a short tRNA-like molecule, mascRNA, and a long MALAT-1 containing a poly(A) tail, which localizes to the cytoplasm, suggesting that they possess distinct activities [31]. MALAT-1 is co-localized to nuclear speckles with the members of serine/arginine-rich (SR) splicing factors, regulating their nuclear localization [32]. Furthermore, MALAT-1 can bind to SR protein and regulate its phosphorylation, which then regulates the alternative splicing of pre-mRNA [33]. MALAT-1 may regulate chemokine receptor pre-mRNA processing by activation of SR protein, thereby affecting chemokine receptor expression in OSCC and impacting its migration. Thus, further studies are required to determine whether the effects of MALAT-1 suppression observed in the present study are mediated through its regulation of SR protein.
In the present study, the expression of UCA1, BC200, and SRA was comparable between OSCC and normal tissues. This is not consistent with previously reported studies describing UCA1 overexpression in bladder cancer [12], BC200 overexpression in lung cancer, breast cancer, cervical cancer and ovarian cancer [15]; and SRA overexpression in breast cancer [16]. This is likely due to specific differences between tumor types.
The present study has some limitations that warrant consideration. Firstly, the sample size used in this study was relatively small, and the correlation between MALAT-1 expression and patient clinical characteristics, including tumor stage recurrence, lymph node metastasis, and patient prognosis, was not analyzed. Furthermore, the impact of MALAT-1 overexpression on SCC4 cells was not determined. Also, the specific regulatory mechanisms by which MALAT-1 silencing altered SCC4 cell growth and migration were not determined, and will be examined in further studies by our group. Finally, MALAT-1 function was only analyzed in a single cell line; thus, further studies in other tongue squamous cell carcinoma cell lines are required to confirm these results.
In conclusion, MALAT-1 is highly expressed in OSCC as compared to adjacent normal tissue. Silencing the MALAT-1 gene inhibited SCC4 cell proliferation, anchorage-independent growth and migration and promoted apoptosis. Thus, MALAT-1 may play an important regulatory role in OSCC progression. Further studies with larger sample sizes are necessary to confirm these findings and correlate MALAT-1 expression with patient outcomes. In addition, studies to elucidate the mechanism through which MALAT-1 expression impacts SSC4 cell growth and migration are necessary.