Introduction
Scabies is a widespread parasitic disease caused by infestation with female mites of Sarcoptes scabiei var. hominis [1, 2]. Two hundred million people are estimated to be affected by scabies worldwide [3]. There are no uniformly agreed diagnostic standards of scabies infestation [3]. Usually, the diagnosis is based on clinical signs, but scabies can mimic other dermatological diseases, which delays diagnosis [4]. Scabies confirmation is based on the detection of Sarcoptes scabiei, its eggs or faeces by microscopic examination of skin scrapings [5]. In a classic infestation, only about 10 to 15 adult female mites are present, which makes microscopic examination a low sensitivity method with a high risk of false-negative results [2].
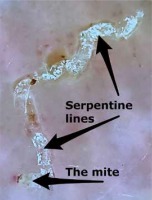
Another method used in scabies diagnosis is dermoscopy [6]. Dermoscopy was originally known as a differentiation tool for the diagnosis of pigmented lesions or melanoma, but it was later proven effective in diagnosing scabies in vivo [7]. Entodermoscopy was a new term proposed in 2006 by Scanni and Bonifazi for dermoscopic diagnosis of ectoparasitosis, which was extended to include superficial skin infections by Zalaudek [8, 9]. Dermoscopy in scabies allows for the identification of triangular or v-shaped, brown structures that correspond to the head and the pairs of mite’s legs, as well as serpentine lines reflecting burrows of scabies (Figure 1). The image resembles a jetliner with its trail, a delta glider or a spermatozoid [6]. In 2019, Scanni proposed a new concept for describing scabies through entodermoscopy: the so-called Mite – Gallery Unit (MGU) [10]. The Mite – Gallery Unit is divided into three parts: the Head with the mite, the Body representing the burrow containing eggs and faeces and the Tail at the end of the tunnel. The MGU allows dermatologists to recognize tunnels without Sarcoptes or with signs of severe inflammation. Furthermore, confocal reflection microscopy and videodermatoscopy are also used in non-invasive scabies diagnosis [11].
In recent years, nucleic acid amplification tests (NAATs) have gained more significance in the diagnosis of parasitic diseases. Polymerase chain reaction (PCR) is a specific tool for the detection and confirmation of Sarcoptes scabiei [5]. Wong et al. [12] developed a conventional and real-time PCR of cytochrome c oxidase subunit 1 (cox1) gene as a useful method for diagnosing scabies. Hahm et al. [13] designed new primers for a nested PCR setup increasing sensitivity. In both studies, skin scrapings were prospectively collected for PCR from patients with suspected scabies. Wong et al. additionally conducted a microscopic examination, while Hahm et al. used DSGSS-ME (DermoScopy-Guided Skin Scraping with Microscopic Examination). Studies demonstrated the usefulness of PCR in diagnosing scabies and its superior sensitivity compared to microscopic examination. There are also studies utilizing non-traumatic skin swab specimens, which need further evaluation [5, 12].
Aim
Diagnostics of S. scabiei infestation is still a significant problem in contemporary medicine as there are no internationally agreed standards for it. The aim of the present study was to compare diagnostic methods for scabies: light microscopy examination of skin scrapings, dermoscopy and real-time PCR testing of skin scrapings. Additionally, we compared the effectiveness of using skin swabs for real-time PCR with epidermal scrapings real-time PCR and light microscopy.
Material and methods
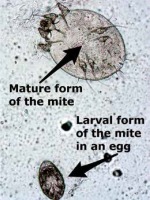
The study was performed at the Dermatology Clinic of the Poznan University of Medical Sciences between March 2017 and February 2019 and involved patients clinically suspected of scabies, such as severe itching, especially aggravated at night, characteristic burrows, papules, nodules, sometimes vesicles or pustules and similar symptoms in other family members. Informed consent was obtained from patients before the examination. Skin scrapings samples were collected after prior dermoscopic evaluation. Patients were examined with a hand-held Delta 20 Plus (Heine Optotechnik GmbH, Germany) dermoscope with a 10× magnification. When a jetliner-like structure was demonstrated, the examination was positive for scabies (Figure 1). Parts of skin scrapings samples were assessed by light microscopy, while the rest was used for real-time PCR testing for Sarcoptes scabiei. Patients were diagnosed with definite scabies when at least one mite or egg was seen under a light microscope (Figure 2). Additionally, some swabs were collected from skin lesions, which were then used for real-time PCR analysis. Standardized sampling was performed using a wet cotton swab (PBS), which was systematically rubbed across suspected lesions.
The research was approved by the Bioethics Committee at the Poznan Medical University of Medical Sciences on 22 June 2017, by Resolution No. 682/17.
DNA isolation
The samples were scraped to a 1.5 ml tube and stored at –80°C prior to isolation. Similarly, swabs were stored at –80°C. DNA was isolated with the use of a QIAamp DNA Mini kit (51304, Qiagen, Germany). Scrapings were put into 160 μl proprietary ATL buffer, 20 μl 1M DTT and 20 μl proteinase K, vortexed for 20 s and incubated overnight at 56°C. Cotton swabs were put to a 2.0 ml tube with 190 μl ATL buffer, 40 μl 1 M DTT and 20 μl proteinase K, vortexed for 20 s and incubated overnight at 56°C. Further steps were followed according to the manufacturer’s instructions. DNA was eluted with 50 μl proprietary AE buffer.
Real-time PCR
The real-time PCR employed here targeted a 120 bp fragment of the cox1 gene of S. scabiei. The primers were the same as in the work of Wong et al. (i.e. primer-1 5’-TGCTATGATTTCTATTGCAACTTTAGG-3’, primer-2 5’-GGGACAGCGATAATTATAGTAGCTGAA-3’) [12], yet we introduced some modifications to the probe. First, the initial sequencing of the amplicon from our first collected isolate marked a substitution mutation at the 10th nucleotide of the probe (GenBank accession MT080809), hence a Y-wobble was introduced. This would increase sensitivity provided that some further isolates showed that variation. Secondly, because the melting temperature of the original probe seemed low, we increased it by elongating the probe by one nucleotide and modifying it with Locked Nucleic Acid (LNA) since there was no possibility to displace the probe to a higher melting temperature region (probe 5’-6FAM-TGTATGAGCYCATCATATATTTACTGTT+GG-BBQ-3’, [+] sign marks the LNA nucleotide). Primers and probes were provided by TIB Molbiol (Germany).
Twenty-microliter real-time PCR reactions were performed with the use of Roche LightCycler 2.0 thermal cycler and Fast Probe qPCR Master Mix (E0422-02, EURx, Poland), including a uracil-N-glycosylase (UNG) treatment for prevention of detecting carried-over amplicons from earlier PCR batches. The ingredients are provided in Table 1.
Table 1
Composition of real-time PCR reactions
The cycling program was as follows: 1) UNG treatment: 37°C, 2 min; 2) predenaturation: 95°C, 3 min, 3) amplification x60: denaturation: 95°C, 10s, ramp-rate (RR) 20°C/s, annealing/elongation: 62°C, 30 s, RR 20°C/s, signal acquisition: single, 530 nm channel; 4) cooling: 40°C. Crossing-points were compared in the absolute analysis mode to a 10-fold dilution standard curve of the aforementioned cox1 amplicon. The standard curve was generated with the same reaction parameters (range: approximately 100k copies to 10 copies). Separate real-time PCR batches were cross-calibrated. All real-time PCR reactions performed were also separated in agarose gel for amplicon confirmation as a precaution (data not shown).
Questionnaire
Patients were asked to complete a questionnaire of our authorship relating to age, place of residence, education, sex, body weight and height, domestic animals, presence, and duration of skin lesions and pruritus among other household members, and treatments used. To evaluate pruritus we used a four-item itch questionnaire by Reich, Mędrek and Szepietowski [14] (Table 2). We also assessed the body surface occupied by skin lesions using the body surface area (BSA) score, a commonly used measure of severity of skin diseases, assuming the rule of nines (9% for head and neck, each arm, thorax, abdomen, upper back, lower back, each thigh, each crus, and 1% for external genitalia).
Results
We have included a total of 75 individual subjects, collected from March 2017 to June 2019. All samples were assessed by light microscopy and all patients underwent dermoscopy examination. So far, all 75 samples of skin scrapings and 41 wet swabs from skin lesions were examined by real-time PCR, while 53 patients agreed to complete the survey. The mean age of the patients was 61.1 years old (range: 8 to 96; median: 63.0) with a female-to-male ratio of 1 : 1.14. The mean age of men was 60.2 (median: 66.0), while the mean age of women was 62.0 (median: 63.0).
Among these 75 samples, 5 positive results (6.67%) were obtained by light microscopy, 8 by dermoscopy (10.7%, including all 5 light microscopy samples), and 21 positive results (28.0%) by real-time PCR from scrapings. Importantly, all light microscopy-positive scrapings were also positive by real-time PCR and all 8 dermoscopy-positive patients were also positive by real-time PCR (Table 3). What is worth noticing is the fact that light microscopy-positive patients with severe scabies (i.e. crusted scabies) also had strong positive results in real-time PCR.
Table 3
Detection of Sarcoptes scabiei, correlations with pruritus and BSA
Out of 41 samples, which were examined by real-time PCR of both skin scrapings and swabs, a total of 15 samples were found positive by any or both (36.6%). In that group only 4 were found positive by both methods, 6 were positive from scrapings but not swabs, and 5 from swabs but not scrapings.
There was no statistically significant correlation between copy numbers of S. scabiei from scrapings and the severity of pruritus as assessed by the aforementioned questionnaire (p = 0.518, rs = 0.100, n = 44). However, if we consider only PCR–positive scrapings, there was a significant correlation between the number of copies and the severity of pruritus (p = 0.023, rs = 0.673, n = 11). Furthermore, there was a weak correlation between copy numbers of S. scabiei from skin swabs and the severity of pruritus (p = 0.07, rs = 0.40, n = 22). If we consider only PCR-positive results, still there seemed to be no statistically significant correlation (p = 0.144, rs = 0.696, n = 6).
There was a statistically significant correlation between copy numbers of S. scabiei from skin swabs and body surface occupied by skin lesions (p = 0.002, rs = 0.626, n = 22). Surprisingly, if we consider only PCR-positive results, there was no statistically significant correlation (p = 0.420, rs = 0.371, n = 6). There was a weak, yet statistically insignificant, correlation between copy numbers of S. scabiei from scrapings and the BSA (p = 0.186, rs = 0.203, n = 44). Although, if we consider specimens with positive results only, the correlation was strong (p = 0.002, rs = 0.825, n = 11).
Finally, we found a statistically significant correlation between copy numbers of S. scabiei from scrapings and the age (p = 0.038, rs = 0.240, n = 75). If we consider only PCR-positive results, this correlation was even stronger (p = 0.018, rs = 0.510, n = 21). The mean age of patients with PCR positive scrapings was 68.0 years old. There was a statistically significant correlation between copy numbers of S. scabiei from skin swabs and age (p = 0.019, rs = 0.366, n = 41). If we consider only PCR–positive results, surprisingly there was no statistically significant correlation (p = 0.385, rs = 0.317, n = 9). The mean age of patients with PCR positive swabs was 75.4.
Discussion
Diagnosis and treatment of parasitic skin diseases have been an important part of medicine for centuries. This is mainly due to the ease of spreading such infections in the surroundings, including the family, schools, kindergartens, hospitals or nursing houses. Scabies is an infectious skin disease, which causes significant morbidity, affects daily functioning and may, through secondary bacterial skin infection like impetigo, lead to serious complications [15]. The clinical features of scabies can mimic other skin conditions, which significantly complicates and delays diagnosis. One can find cases of scabies diagnosed as urticaria, herpetic dermatitis, mycosis fungoides, contact dermatitis or even secondary T-cell lymphoma described in the literature [16]. Data show that 45% of patients with scabies are incorrectly diagnosed [17]. Nowadays, there are no consensus guidelines for scabies, which hinders the interpretation of existing epidemiological and treatment studies [2]. The need for diagnosis standardization of scabies was considered as a priority at the inaugural meeting of the International Alliance for Control of Scabies in Atlanta in 2012 [18]. Scabies is confirmed, when its eggs, mites or faeces are detected under a light microscope. However, the low sensitivity of microscopic examination and a high risk of false-negative results lead to scabies detection rates varying from 10% to 70% [19]. Improvements in scabies detection are therefore of crucial importance.
Another available scabies detection method is dermoscopy. Dermoscopy and reflective confocal microscopy are important, non-invasive methods in the scabies diagnosis, being more patient-friendly than the traditional scraping test [11, 20]. Studies prove that the sensitivity of a manual dermoscope test (with 10× magnification) in the diagnosis of scabies is higher than light microscope examination [4, 19]. Dupuy et al. [20] conducted a study to compare dermoscopy and microscopic examination of skin scrapings in scabies diagnosis. Three dermoscopists were involved: one expert and two inexperienced. Sensitivities in scabies diagnosis were 91% for dermoscopy and 90% for microscopic examination of skin scrapings. Specificities were 86% for dermoscopy and 100% for microscopic examination. Dermoscopy sensitivities were similar for inexperienced dermoscopists and the expert. Furthermore, diagnostic accuracy of inexperienced researchers increased during the study. Even if one of the limitations in using dermoscopy in the scabies diagnosis is the long training period required to distinguish artefacts from positive cases [6], this study shows that dermoscopy can be a useful tool even among inexperienced scientists. According to our study, dermoscopy was more effective than microscopic examination. Among 75 patients, 8 (10.7%) were positive in dermoscopy and only 5 (6.7%) patients were positive in microscopic examination. Importantly, all these 5 light microscopy positive patients were also positive in dermoscopy, indicating that the former might not have been necessary. On the other hand, two dermoscopy results were questionable under light microscopy since it was difficult to distinguish artefacts from positive results. These two were negative in real-time PCR. Therefore, the superiority of dermoscopy over light microscopy requires further examination, yet it seems to be a better screening tool out of these two.
There have already been reports on PCR in scabies diagnosis being more sensitive than a direct scraping examination and light microscopy [12, 13, 21, 22].
In their research, Hahm et al. [13] showed the advantage of nested PCR over conventional PCR and traditional methods. Out of the 63 patients, scabies was diagnosed in 28 patients (44.4%, using DSGSS-ME method), while a positive nested PCR test was obtained in 37 patients (58.7%, additional 9 compared to DSGSS-ME).
Wong et al. [12] examined skin scrapings of 100 patients with suspicion of scabies using microscopic examination and PCR (conventional and real-time PCR, both targeting cox1). Additionally, for 1 patient with crusted scabies, skin swabs were taken for PCR. Sarcoptes scabiei var. hominis was detected in 29 skin scraping samples by PCR (29.0%) and only in 17 by microscopic examination (17.0%). All light microscopy-positive results were also positive in PCR, which indicated high sensitivity of PCR. Skin swabs from the patient with crusted scabies were positive by conventional and real-time PCR. The study demonstrated that mite DNA disappeared within 2 weeks after antiparasitic treatment, which suggests that quantitative PCR might be used as a tool in assessing the outcomes of treatment.
In our study, similar to the majority of others, skin scrapings PCR and dermoscopy were more sensitive in scabies diagnosis than light microscopy examination (28.0%, 10.7%, and 6.7%, respectively). Our real-time PCR and scraping sampling protocols were almost identical to that of Wong et al. and these real-time PCR results are in fact quite similar. Moreover, we have also tested 2 patients with crusted scabies during their treatment. We have found these samples to be positive by real-time PCR after a few days. Other studies [5, 23] also show that PCR can be used as follow-up because of residual scabies material in the epidermis after treatment.
On the other hand, there have been claims that PCR might not be that effective. Delaunay et al. [5] conducted a study to evaluate the use of PCR with only dry swabs as a first-line tool in the diagnosis of scabies when an epidemic is suspected. Out of 164 patients, scabies was diagnosed in 87 people (by dermoscopy or clinical examination), while positive PCR was obtained in only 33 patients. Standardized dry swab PCR reached 20.1%. The authors proposed a study protocol, in which if at least 1 patient in a group has a positive PCR result, then the whole group requires treatment. If all PCR results are negative, then expert judgment is required. Based on that it seems reasonable to treat the entire household if at least one member is found positive by swab-based PCR.
Considering swab sampling only, our results were not much better. We identified 9 positive swabs out of 41 (22.0%) of suspected patients, although the swabbing and PCR techniques were greatly different. Swabs taken by Delaunay et al. were dry, ours were wet. Their target was chromosomal (ITS2) [5], ours was mitochondrial. It seems possible to boost sensitivity higher through technological optimization. If that is in fact achievable, swabbing could replace other sampling techniques. Yet, this claim requires further examination. For the moment though, based on our preliminary results, we recommend taking both scrapings and swabs for the highest PCR sensitivity.
All of the aforementioned sensitivity values of PCR appear low. This is because PCR positive samples are compared to the initial number of patients chosen for the analysis. One has to keep in mind however that these initial visual symptomatic inclusion criteria should not be considered gold standard, nor any standard whatsoever, as not only do they apparently generate a significant portion of false positives, but they also differ greatly in between facilities. In fact, there is no uniformly accepted gold standard.
The positive correlation between the number of copies from real-time PCR-positive scrapings and the severity of pruritus confirms that pruritus is an important symptom in patients with already diagnosed scabies. We can conclude that pruritus is obviously not a certain criterion as it correlates with these PCR positive patients, but not the whole studied group.
Similarly, we found a strong positive correlation between the number of copies from real-time PCR-positive scrapings and the severity of skin lesions, while there was no correlation between the severity of skin lesions and real-time PCR results from both positive and negative patients together, suggesting that clinical signs cannot be an unambiguous criterion for positive diagnosis of scabies.
Interestingly, contrary to scrapings, wet swab-based real-time PCR seems to somewhat correlate with pruritus (p = 0.07) and BSA (p = 0.02) in the whole group but perhaps not that much in positive samples. However, these results should be treated with caution due to low positive sample groups. This indicates that a swab-based NAAT could be superior in the initial diagnosis of scabies, provided that the sensitivity levels are higher, while molecular analysis of scrapings, on the other hand, might be used e.g. in treatment assessment. Again, for the moment we recommend doing both.
The positive correlation between copy numbers from scrapings and skin swabs with age confirms that the frequency of scabies increases with age. According to Kim and Cheong [24], who presented the annual and seasonal trends of prevalence of scabies in South Korea, the annual prevalence increases with age and peaks at 3.0–4.1 per 1000 people in the elderly over 80 years old.
The correlation between the copy numbers from skin swabs and age was statistically significant in the whole group, while not statistically significant in positive samples only. The latter was probably the effect of a small study group (9 real-time PCR positive skin swabs, yet not all of the patients completed the survey).
Studies of the S. scabiei genome are considered to be a great opportunity for the development of prevention, early diagnosis and effective treatment. Next-generation sequencing (NGS) allows studies on all skin microbiomes, including bacteria, fungi, viruses, and mites, by providing many sequences from a sample [25, 26]. Based on molecular analysis, scabies in humans are caused by four S. scabiei groups (varieties) [26]. The size of the genome, based on quantitative PCR analysis, had been estimated at about 96 Mbp and the number of chromosomes – 17 or 18. The genome of scabies seemed about twice as small as the genome of the house mite, which is believed to be an adaptation to the parasitic lifestyle [27]. A later NGS paper reports a draft genome size of as low as 56 Mbp [26].
Conclusions
Although real-time PCR requires special equipment, it is more expensive and time-consuming than light microscopy or dermoscopy, it can identify scabies at an early stage of infestation when the number of parasites is still low. Therefore, the risk of scabies outbreaks can be minimized.
At the moment it seems reasonable to suggest performing two separate real-time PCRs from both scrapings and swabs. We have not analysed the effect of combining these samples prior to or after DNA isolation to perform just one reaction. Nevertheless, we would suggest further studies to be targeted at improving swabbing and PCR techniques, to possibly eliminate the need for taking scrapings as swabs are far less invasive and safer for the medical personnel. The utility of using PCR as an alternative method for scabies diagnosis is promising, yet needs further evaluation.