Introduction
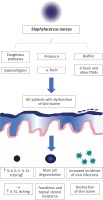
The pathogenesis of atopic dermatitis (AD) is complex and remains unclear. It develops in subjects with a genetic predisposition under the influence of different environmental factors [1, 2]. The skin microbiome and its fluctuations are directly associated with this disease. This review mainly focuses on the role of Staphylococcus aureus in the development and perpetuation of AD (Figure 1).
Microbiome – host interactions
Numerous species of bacteria, viruses and fungi colonize human skin and are collectively defined as its microbiome [3, 4]. To understand the role of the skin microflora in the pathogenesis of AD, it is necessary to explore its qualitative and quantitative composition. The advent of Next Generation Sequencing (NGS) has facilitated the study on human microbiota. Grice et al. described the bacterial profile of 20 areas of the human skin thanks to the analysis of 16S ribosomal RNA (16S rRNA) [5]. A total of 19 phyla were distinguished, four of which were dominant: Actinobacteria (59% of total bacterial species, genera of this phylum – Micrococcus, Propionibacterium, Corynebacteria), Firmicutes (24%, Lactobacillus, Streptococcus, Staphylococcus), Proteobacteria (17%, Paracoccus, Hematobacter) and Bacteroidetes (7%, Prevotella, Porphyromonas). As expected, each microorganism was isolated from niches where local conditions (moisture level, pH, lipid profile etc.) favoured its growth. Regardless of these differences, it must be pointed out that the healthy skin harbours extremely diverse microflora.
Similarly to gut microbiome, it has been reported that skin commensal bacteria influence many important processes, including the maturation of the immune system [3]. The analysis of prevalence of atopic diseases in children, who were raised in different environments (rural vs. municipal) led to the elaboration of the “hygiene hypothesis” [6, 7], according to which insufficient exposure to antigens of diverse microorganisms may lead to excessive Th2-type immune response. Cytokines produced by Th2 lymphocytes (interleukin (IL)-4, -5, -13) are strategic players in the vicious cycle of atopic diseases. They are also known to aggravate AD through many mechanisms [8–10].
Non-altered microbiome helps in maintaining skin homeostasis. Naik et al. have proven that skin commensals provide a specific “training” for the immunological system [11]. Their presence results in increased signalling through IL-1R which inhibits Th2 polarization and ensures correct immunological response to pathogenic microorganisms. Furthermore, specific bacteria are capable of inhibiting the growth of potentially harmful microorganisms e.g. by producing bacteriocins and preventing biofilm formation. These mechanisms will be presented below in more detail.
Genetic disorders hampering the control of bacterial pathogens
Pattern recognition receptors (PRR) are a conservative element of the innate immune system. They bind pathogen-associated molecular patterns (PAMP) and danger-associated molecular patterns (DAMP) [12]. PRR-mediated signalling leads to nuclear factor-kB (NF-kB) activation which results in production of pro-inflammatory cytokines and anti-microbial peptides (AMP). Toll-like receptors (TLR) are a PRR subtype, which orchestrate an appropriate immune response. Some reports have shown down-regulation of TLR2 (S. aureus-antigen binding receptor) in keratinocytes and monocytes of patients with AD [13, 14]. Reduced TLR2 expression has been associated with decreased production of IL-1B, tumor necrosis factor-α (TNF-α) and IL-8 by monocytes [14]. Additionally, TLR2 seems to be important for maintaining the skin barrier function as S. aureus antigens and synthetic TLR2 agonists have been shown to stimulate the production of tight-junction proteins (claudin-1 and claudin-23) [15]. Of note, patients with short-term S. aureus colonization tend to have an increased expression of claudin-1 and -4, while prolonged presence of this pathogen on the skin down-regulates their expression and negatively affects the barrier function provided by tight-junctions [16]. Clinical importance of these observations remains unclear.
Destruction of the epithelial barrier and increased skin permeability are another prominent characteristics of AD which predispose to bacterial colonization. The patients with atopic dermatitis show deficiencies in two crucial components of the skin barrier: ceramides and intracellular connections [17–20]. While they are not directly related to negative microbiome fluctuations, they facilitate them for numerous reasons:
Epithelium damage leads to the increased expression of adhesive molecules [21–23]. This, on the other hand, favours adherence of pathogenic bacteria to the skin.
Increased skin permeability enables local penetration of allergens and irritants. They induce in-situ inflammation which further debilitates immune responses and epithelial regeneration [24, 25].
Trans-epidermal water loss (TWLE) and skin dryness are increased, which combined with local inflammation causes pruritus and the secondary barrier damage due to scratching [26].
Recent genetic analyses performed by Japanese and German scientists have proven the existence of numerous polymorphisms predisposing to AD, some of which are related to the immune system functioning [27, 28]. Nevertheless, some data suggest that investigation should not be limited to coding sequences of the human genome only. Gene expression may also be deficient because of epigenetic disorders or interference of miRNA in the process of translation. miRNAs have been reported to regulate the expression of TLRs and influence mechanisms of responses to superantigens. These observations may justify the attempts of targeting them to reduce AD symptoms [29–31].
Microbiome characteristics in atopic dermatitis
Many diseases alter natural composition of the microbiome. Two well-known examples include selective expansion of Clostridium difficile in antibiotic-associated diarrhoea [32] or excessive development of diverse anaerobic microflora with simultaneous reduction in Lactobacilli in bacterial vaginosis [33]. Atopic dermatitis is characterized by specific changes of the microbiome as well.
Classic culture methods have revealed that skin lesions in AD are colonized by S. aureus in up to 90% of patients and that this effect is the most prominent in samples collected during disease exacerbation [34, 35]. These observations have been confirmed by the analysis of bacterial 16S rRNA [36]. Staphylococcus aureus is associated with the reduction in total microbiome diversity which positively correlates with disease severity. Therapeutic interventions that restore natural skin microflora are known to alleviate the symptoms of AD [37, 38]. Recent investigations in children imply that colonization of the skin by S. aureus may not only be an exacerbating factor of AD, but also lead to its development [39]. Nonetheless, infants whose gut was colonized by S. aureus strains carrying a specific combination of superantigen and adhesin genes showed a reduced risk of subsequent development of atopic eczema, which suggests that the presence of this pathogen may globally promote appropriate maturation of the immune system [40].
Much effort has also been dedicated to determining the role of S. epidermidis in AD. Unlike S. aureus, colonization by commensal staphylococci (e.g. S. epidermidis) at the age of 2 months was associated with a decreased risk of AD development at the age of one year [41]. Although S. aureus expansion tends to inhibit the growth of most skin commensals, S. epidermidis remains relatively widespread in AD lesions [42]. Hon et al. have shown that aggravation of AD symptoms and increased colonization of the lesional skin by S. epidermidis are associated [43]. Nevertheless, they have also reported that S. epidermidis inhibits S. aureus growth in the lesions. This phenomenon could be explained by the ability of S. epidermidis to stimulate AMP production through TLR2 signalling [44] and by secretion of phenol-soluble modulins (PSM) and bacteriocins that act synergically with the aforementioned in controlling pathogens such as S. aureus and S. pyogenes [45, 46]. Furthermore, S. epidermidis can inhibit formation of biofilm by S. aureus, while lipoteichoic acid of this specie shows immunomodulatory properties in healing wounds [47–50]. The latter stresses the importance of natural commensals in proper skin recovery and should draw the attention of future research in the context of AD lesions.
Staphylococcus aureus – virulence factors affecting the disease course in atopic dermatitis
Staphylococcus aureus not only causes skin infections, but also influences the course of AD by seemingly asymptomatic colonization. This effect can be explained by its ability to produce virulence factors such as biofilm, superantigens, α-toxin and protein A that are considered as important elements of the vicious cycle of AD.
Biofilm
Some strains of S. aureus are capable of biofilm formation. While this process is commonly associated with decreased effectiveness of antibiotic treatment, it may also have other consequences. Biofilm impedes clearance of pathogenic bacteria by shielding them from host immune cells such as neutrophils and macrophages. It is also known to facilitate macrophage cytotoxicity [51]. Furthermore, apoptosis has been observed after exposure of keratinocytes to staphylococcal biofilm in vitro, and substances released upon this process (TSLP, IL-4 and IL-13) showed a negative influence on the clearance of pathogenic bacteria and skin regeneration [51]. Chronic skin damage associated with S. aureus colonization creates a favourable environment for biofilm development [52].
Exogenous proteases
Epithelial barrier integrity may be impaired by exogenous proteases of S. aureus which eases the penetration of environmental antigens into deeper compartments of the skin [53, 54]. It makes the lesions of patients with AD prone to exacerbation in the presence of allergic substances and other irritants. Additionally, recent studies have focused on the ability of S. aureus to penetrate into sub-epidermal skin layers. Nakatsuji et al. have found that this process depends on the viability of S. aureus strains and their ability to produce proteases [55]. Presence of S. aureus in the dermis has been correlated with an increased expression of IL-4, IL-13, IL-22 and TSLP as well as with a decreased production of cathelicidin [55]. Another study suggests that SspA/V8 protease is the main substance compromising the epithelial barrier [56]. This process is prevented by IL-1B production and subsequent secretion of human b-defensin 2 (hBD2), giving light to elaborate novel therapies. Moreover, Sonesson et al. have discovered that S. aureus can produce another protease group, staphopains, which inactivate AMPs (LL-37) responsible for the degradation of bacterial biofilm [52].
Staphylococcal superantigens
Some strains of S. aureus produce superantigens. The most important ones include staphylococcal enterotoxin A and B (SEA/B) and the toxic shock syndrome toxin-1 (TSST-1) that are known to affect the course of AD in a negative way [57–65]. These molecules are bound by local antigen presenting cells (APCs) and presented to other elements of the immune system via major histocompatibility complex class 2 (MHC-II) [64]. They cause a chaotic inflammatory response by activating a large percentage of naïve T lymphocytes, which leads to excessive Th2 cytokine release [57, 60]. It has also been demonstrated that despite their regulatory phenotype, CD4+ FOXP3 cells present in the skin of patients with AD can be stimulated to secrete Th2 cytokines after exposure to staphylococcal superantigens [66]. Furthermore, SEB (but also other staphylococcal antigens, such as lipoteichoic acid) has been associated with increased synthesis of IgE [64, 65, 67]. This supports the theory that the presence of S. aureus on the skin can provoke AD flares instead of being just an exacerbating factor. In the study conducted by Hon et al. [68], SEB-specific IgE concentration has been positively correlated with disease severity. Another negative function of SEB includes its availability to induce monocyte apoptosis by up-regulating TNF-α [69]. It has also been reported that S. aureus superantigens can contribute to the effect of glucocorticoid and tacrolimus resistance, which is especially troubling from the therapeutic point of view [62, 63].
Selected staphylococcal superantigens are also involved in the pathogenesis of itch in AD. IL-31, a molecule whose levels have been correlated with the intensity of pruritus, has been shown to be up-regulated after exposure to staphylococcal superantigens in vivo and more particularly to SEB (in vitro) [70]. Studies have shown that increased levels of IL-31 negatively affect the production of AMPs, which impedes S. aureus clearance [71]. What is more, skin damage and keratinocyte necrosis caused by constant scratching can additionally lead to sensitization to autoantigens, which drives the vicious cycle of AD.
α-Toxin
Staphylococcal α-toxin (also known as α-hemolysin) is highly cytotoxic to keratinocytes by leading to intrinsic caspase activation and cytochrome c release from the mitochondria in experimental studies [72]. It has been demonstrated that keratinocytes exposed to Th2 cytokines are more susceptible to apoptosis through this mechanism [73]. Hong et al. have shown that α-toxin induces atopic dermatitis-like skin inflammation and disruption of the skin barrier [74]. α-Toxin production by S. aureus positively correlates with disease severity and has been proposed as a potential diagnostic and therapeutic target for the control of AD. Moreover, α-hemolysin is upregulated upon formation of biofilm following neutrophil exposure and causes neutrophil inhibition. Interestingly, it also facilitates viral entry into the cells. It may be one of the reasons why patients with AD are predisposed to infections with pathogens such as Molluscum contagiosum or Herpesviridae (increased incidence of eczema herpeticum) [42, 72, 75].
δ-Toxin and other phenol-soluble modulins (PSMs)
Phenol-soluble modulins (PSMs) are a family of short peptides produced by staphylococci [76]. PSMs secreted by S. aureus show cytolytic activity in leukocytes and are thought to play an important role in the pathogenesis of AD. δ-Toxin of S. aureus is one of the PSMs investigated in this context. The study conducted by Nakamura et al. [77] has revealed that exposure of mast cells to δ-toxin leads to their degranulation rather than lysis. Excessive IL-4 and IgE production with simultaneous exacerbation of Th2-mediated inflammation in the skin has been observed as well [77]. The PSMs have also been shown to cause keratinocyte lysis with subsequent release of proinflammatory cytokines such as IL-18 and IL-1β [78].
Staphylococcal protein A
Staphylococcal protein A is constitutively expressed on the surface of S. aureus and contributes to evasion of host immune responses [79]. Jun et al. have reported that it is highly expressed in membrane vesicles of S. aureus, whose application to AD-like skin lesions induces eczematous dermatitis. Based on this observation, the authors suggested that S. aureus may be an inducing factor in AD. Another study has revealed that protein A along with toxins of S. aureus induce local inflammation mediated by TNF-α [80].
Novel therapeutic methods with the aim of restoring natural composition of the skin microbiome
Increasing knowledge about the significance of disordered skin microbiome in AD is prompting the search for new treatment methods. Routine management of patients with AD with antibiotics would be contraindicated mainly because of the extent and chronic course of disease, cost-effectiveness, selection of drug-resistant strains of pathogenic bacteria [81, 82] and reduction of commensal microflora. In recent years many ground-breaking strategies have been developed. For example, lysate of a recently discovered Gram-thermal spring bacterium called Vitreoscilla filiformis has been reported to significantly reduce the symptoms of AD and seborrheic dermatitis [83–85]. Volz et al. [86] have revealed that V. filiformis antigens induce IL10+ dendritic cells. They are a source of stimuli responsible for naïve CD4+ lymphocytes maturation into Treg1 lymphocytes. Treg1 lymphocytes produce IL-10, a cytokine known for its strongly immunosuppressive properties. This process depends on TLR2 signalling, which has already been mentioned in the context of AD-specific disorders of the innate immune system [87]. Another trial has revealed that application of lotion with heat-treated probiotic strain Lactobacillus johnsonii for 3 weeks controlled S. aureus colonization and reduced disease severity measured with the SCORAD index [88]. Interestingly, a higher concentration of S. aureus at baseline was associated with good response to the treatment. Nakatsuji et al. [89] have analysed the relation between the presence of commensal coagulase-negative staphylococci (CoNS) S. epidermidis and S. hominis and skin colonization by S. aureus. They have found that CoNS produce lantibiotics, i.e. prokaryotic antimicrobials that along with human AMPs (LL-37) lead to the clearance of S. aureus in animal and human models [89]. Favourable results have also been reported for topical microbiome transplantation with Roseomonas mucosa [90].
Other trials have focused on a potential therapeutic effect of synthetic AMPs. In the study conducted by Dawgul et al. [91], citropin 1.1 and temporin A appeared to be active against all S. aureus strains. Furthermore, unlike conventional antimicrobials, they did not induce resistance and showed anti-biofilm activity. The study of several AMPs introduced in catheter infections caused by biofilm-forming S. aureus has revealed high effectiveness of the treatment, especially in case of D-Bac8c2,5Leu [92]. Additionally, some promising data have come from the research over all-D omiganan, which has shown bactericidal activity towards S. aureus and satisfactory half-life in vivo [93].
Another promising direction of research includes phagolysins, i.e. anti-infectives derived from bacteriophages which have been shown to resensitize bacteria to antibiotics [94, 95]. Totté et al. [96] published a case series reporting successful treatment of MRSA infections with Endolysin Staphefekt SA.100. Results of a multi-centre randomized trial designed to evaluate the influence of Staphefekt on the use of corticosteroids, disease severity, quality of life and composition of the microbiome in patients with AD are to be expected shortly [97]. Finally, Baldry et al. have managed to block production of α-toxin by inhibiting agr signalling and have subsequently obtained reduction of inflammation in an animal model [98]. Nevertheless, an additional studies to verify the clinical application of these novel therapies in AD is needed.
Conclusions
Despite its complexity, pathogenesis of atopic dermatitis is quite obviously associated with the skin microbiome. The role of S. aureus in the vicious cycle of AD seems to be exceptionally pronounced. S. aureus eliminates commensal bacteria from the skin, while its virulence factors show a negative effect on the epithelial barrier integrity and immune system functioning. It is possible that S. aureus is not only a secondary exacerbating factor but it is also one of the reasons why AD flares occur. Physiologically, the microbiome is responsible for the maintenance of immunological homeostasis and reduction of skin colonization by pathogenic bacteria. Therefore, therapies aimed to restore natural composition of the skin microbiome are being elaborated.