Purpose
Cervical cancer is a common malignant tumor in women [1]. External beam radiotherapy combined with brachytherapy is the standard radiotherapy treatment for cervical cancer. At present, image-based three-dimensional (3D) brachytherapy has become the standard procedure. Applicator reconstruction is a critical step in treatment planning [2]. At this stage, the applicator reconstruction is performed manually by the planner.
Automatic planning is an important research topic in radiotherapy [3]. Automatic, accurate, and rapid applicator reconstruction needs to be resolved for automatic planning in brachytherapy [4]. Before deep learning, researchers usually used threshold-based method to segment the applicator; however, this method still requires planners to define some points manually in clinical practice [5]. In recent years, more studies on automatic applicator reconstruction have been conducted based on deep learning [6-12].
In this study, a deep learning model to automatically segment and reconstruct the applicator was built. The dosimetric differences were compared between manual reconstruction and automatic reconstruction.
Material and methods
Segmentation model
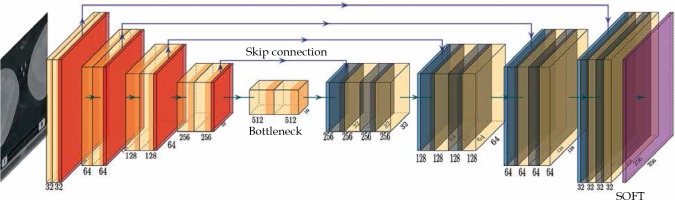
The network structure is illustrated in Figure 1. The model was based on a U-Net structure and consisted of 10 components [13]. The model input included two-dimensional (2D) computed tomography (CT) images; the first 5 layers were under sampling layers, each layer contained two convolution operations and max pooling operation. Layers 6 to 9 were up sampling layers. After the transposed convolution operation, each layer merged the low-level information with the high-level information through a skip connection and then, performed two convolution operations. The convolution kernel was 3 × 3, the transposed convolution kernel and the max pooling kernel were 2 × 2, the step was both 1, and the activation function was Relu. The 10th layer integrated the cross-channel features through a 1 × 1 convolution layer, the activation function was sigmoid, and finally derived the desired 256 × 256 2D mask image.
Data annotation
We retrospectively studied 70 patients who completed CT-based 3D brachytherapy. All patients used a CT/magnetic resonance imaging (MRI) Fletcher applicator (Elekta part # 189.730). The resolution of CT images was 1 mm × 1 mm, and the slice thickness was 3 mm. The number of CT slices was 69 to 100 (average, 87). The 70 patients were divided into training data, validation data, and test data according to the ratio of 50 : 10 : 10. The annotating of applicator was performed by an experienced physicist using Oncentra (Elekta AB, Stockholm, Sweden, version 4.3) treatment planning system. The tandem diameter of CT/MRI Fletcher applicator was 4 mm, and the inner lumen of tandem was about 2 mm, so the applicator from the middle of each channel was annotated and depicted in a circle with a 2 mm radius. The number of CT slices containing mask images (ground truth) was from 38 to 71, and the average number of slices for training data and validation data was 58 and 56, respectively.
Data pre-process
We performed historical equalization on the CT images in the training data and validation data to raise the applicator characteristics. For cervical cancer patients, the applicator is usually located in the middle area of CT image. In order to reduce the training data and validation data size, we shortened the CT images to a small region that included the Fletcher applicator (256 × 256 pixels). The shorten center was the geometric center of the CT image. We normalized all CT images and ground truths, so that all data were between 0 and 1. Deep learning model generally requires a large number of training data to learn effectively and prevent under-fitting or over-fitting. Therefore, we used ImageDataGenerator interface of Keras to augment the training data. More images were generated by rotating, enlarging, scaling, and shifting the image in other directions. The final training data and validation data were 256 × 256 × 116 × 50 and 256 × 256 × 56 × 10, respectively.
Training and segmentation
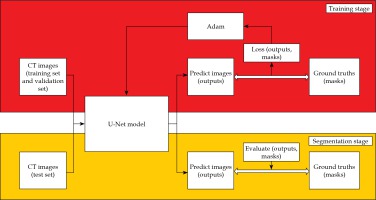
The training and segmentation were completed on an Intel Core i7-7700HQ CPU @ 2.80GHz, GPU NVIDIA GeForce GTX 1050ti, 8GB RAM personal computer, and the process is illustrated in Figure 2. In the training stage, the pre-processed training data and validation data were placed into the U-Net model. The batch size was 8, the epoch was 200, and the initial learning rate was 0.0001. The learning rate was dynamically adjusted by monitoring the learning process, and early stopping was adopted to avoid over-fitting of the model. We selected an Adam (adaptive moment estimation) as the optimizer and a dice loss as the loss function in the training stage [14]. The dice similarity coefficient (DSC) and dice loss were defined as follows:
where A is the prediction mask image, B is the ground truth, and λ is the Laplace smoothing factor (usually 1), which could reduce over-fitting and avoid the denominator, and is 0.
After the training, the data of the test set were inputted into the model for segmentation, and the segmentation result of the corresponding patient was obtained and evaluated.
Evaluation of segmentation results
We used two parameters to evaluate segmentation results [15, 16]. The first one was DSC. The DSC was used to measure the similarity of the two segmentation samples (manual and automatic). The range of DSC was 0-1. The larger the DSC, the better the segment. In the test process, the mean and standard deviation of DSC were calculated for all slices.
The second parameter was Hausdorff distance (HD). The HD was defined as:
where A is the prediction image surface, B is the ground truth surface, and a and b are the points on the surfaces of A and B, respectively. In order to eliminate the influence of outliers between the predicted image and the ground truth, the 95 percentile HD (HD95) was calculated. The unit of HD95 was mm. The smaller the HD95, the better the segmentation.
Automatic applicator reconstruction
We applied the test data into the trained model. The segmented applicator contour was composed by multiple points. We used a clustering method to create applicator contours as showed in reference [12]. For each channel, the average coordinate value of all points in one slice was calculated to obtain the trajectory of the channel central path. A polynomial curve fitting method was used in the reconstruction to reduce the systematic error. Then, the trajectory was written into the RT structure file, and the process of automatic reconstruction was completed.
Evaluation of reconstruction results
For each patient, we used the tip error and the shaft error to evaluate the reconstruction results [17]. The definitions of tip error and shaft error were as follows:
where N is the total channel number (3 in this study), Predi is the predicted length of the i-th channel, and Gti is the annotation length of the i-th channel.
where M is the slices number, Pred(x, y) are the predicted coordinates of the i-th slice, and Gt(x, y) are the annotation coordinates of the i-th slice.
Dosimetric comparison
Dose volume histogram (DVH) parameters were used to evaluate the dosimetric difference between the automatic reconstruction and manual reconstruction. The DVH parameters were D90% for high-risk clinical target volume (HR-CTV), and D2cc for organs at risk (OARs). The OARs included bladder, rectum, sigmoid, and intestines [18].
Results
In the training stage, the loss converted to a lower level after 10 epochs. Because of the early stopping, the model finished training after 60 epochs. At the end of training stage, the loss of training data and validation data decreased to 0.10 and 0.11. The average DSC was 0.90 for the training data and 0.89 for the validation data. The training results indicated that there was no over-fitting. The total training time was 8 hours for 60 epochs.
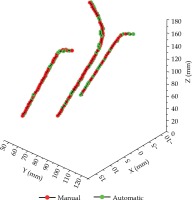
The segmentation and reconstruction results are presented in Table 1. In the average overall test data, the applicator segmented DSC was 0.89 and the HD95 was 1.66 mm. Compared with the manual reconstruction, the average tip error of the 10 cases was 0.80 mm, and the shaft errors were all within 0.50 mm. Tip error and shaft error of three channels were within a reliable range. Table 2 shows the breakdown time in this model. The average total time (including pre-processing, segmentation, and reconstruction) was 17.12 s. A reconstruction comparison of Fletcher applicator is illustrated in Figure 3.
Table 1
The results of applicator segmentation and reconstruction on the test cases
Table 2
Breakdown time (s)
In order to obtain a more conservative result, we chose 1 mm, which doubled the maximum shaft error (0.5 mm), to compare dosimetric differences. Table 3 presents the dosimetric data obtained by the two different reconstruction methods. Although we increased the shaft error, the dosimetric differences of HR-CTV D90% were still less than 0.30%, and the maximum 2.64% for OARs D2cc. These results confirmed that the accuracy of this model was acceptable [19].
Table 3
The results of dosimetric differences between manual and automatic reconstructions
Discussion
Applicator reconstruction is one of the most critical steps in brachytherapy treatment planning [2, 20]. Motivated by recent advances in deep learning, we investigated a deep learning method to automatically segment and reconstruct applicators in CT images for cervix brachytherapy treatment planning with the Fletcher applicator. Evaluation results proved its feasibility and reliability. The model can quickly and accurately segment the applicator regions, and complete the reconstruction. For the commonly used applicator reconstruction, this model takes about 17.12 s from pre-processing to reconstruction, while an experienced physicist needs about 60 s, which increases the reconstruction efficiency by almost 4 times. The pre-processing, segmentation, and reconstruction time would be shorter if a high-performance computer was used.
Many scholars have conducted a series of research on the reconstruction of interstitial needles. Zhang et al. constructed an attention network and applied it to ultrasound-guided high-dose-rate prostate brachytherapy [6]. Wang et al. built two kinds of neural networks for segmentation of interstitial needles in ultrasound-guided prostate brachytherapy [7]. By constructing a 3D U-Net network, Zaffino et al. completed the reconstruction of interstitial needles in MRI-guided cervical cancer brachytherapy [17]. Moreover, Dai et al. developed a deeply supervised model by an attention-gated U-Net, incorporated with total variation regularization to detect multi-interstitial needles in MRI-guided prostate brachytherapy [8].
There are also studies about applicator segmentation. Hrinivich et al. studied an image model algorithm to reconstruct the applicator in cervical cancer guided by MRI. The average reconstruction accuracy of ring applicator and tandem applicator were 0.83 mm and 0.78 mm, respectively [9]. Based on the U-Net, Jung et al. proposed a deep learning-assisted applicators and interstitial needles digitization method for 3D CT image-based brachytherapy. In tandem and ovoid applicator digitization, DSC reached 0.93 and HD was less than 1 mm [10, 11]. Deufel et al. applied image thresholding and density-based clustering in applicator digitization. Their HDs were ≤ 1.0 mm, and the differences for HR-CTV D90%, D95%, and OARs D2cc were less or equal to 1% [12]. In the present study, the DSC was 0.89, HD was 1.66 mm, the dosimetric differences for the target were less than 0.30%, and the maximum 2.64% for OARs D2cc. Compared with previous studies, our results still have room for an improvement.
After the model trained with the Fletcher applicator, six patients with a vaginal CT/MRI applicator (Elekta part # 101.001) were also used to test this model. A vaginal applicator differs from the Fletcher applicator with a connection end. The average overall test data DSC, HD95, tip error, and shaft error were 0.84, 1.81 mm, 1.00 mm, and 0.31 mm, respectively. The dosimetric difference of HR-CTV D90% was less than 0.51%, and the one of OARs D2cc was less than 4.87%. The results of vaginal applicator were slightly worse than that of the Fletcher applicator, since the model was trained by Fletcher applicators; however, the differences of all evaluated parameters were less than 5%.
Automatic radiotherapy planning is a hot spot in current studies, and it is also a subject of interest of our research group. We have made efforts in this direction [21, 22]. These results prove that this model could be integrated into an automatic treatment planning system.
Our present study has some limitations. One is that, according to published research, the accuracy needs to be improved. Although the dosimetric differences between the two reconstruction methods were acceptable, we are still working on the ways to increase the segmentation and reconstruction accuracies. The other limitation is that only two applicator types were included in this study. Here, we chose the Fletcher applicator because it is one of the most commonly used applicators in our center. Even though there are many kinds of applicators in clinical practice, this model could not be used for other applicator types. However, we are convinced that reconstruction of other applicator types can be carried out quickly with the foundation of the present research. Another limitation of this work is that the CT slice thickness was large in this study (3 mm). Slice thickness is a source of the tip uncertainty; therefore, the large slice thickness could be a reason of the large tip error in this study.
Conclusions
In summary, applicator reconstruction is a critical process of treatment planning. We implemented a U-Net model for applicator segmentation and reconstruction in CT-based cervix brachytherapy in this study. The DSC, HD95, tip error, and shaft error were used to evaluate this model. The results demonstrated that our model is clinically attractive. Therefore, this research paves the way for automatic treatment planning in brachytherapy.