Introduction
People affected with allergic disorders such as allergic rhinitis, allergic asthma, allergic dermatitis and food allergy can encounter acute signs and symptoms of the disease within a few minutes of exposure to allergens. Allergic diseases are the main cause of great morbidity and the rise in allergic diseases has become one of the major health problems nowadays [1, 2].
Bronchial asthma is a chronic inflammatory disease affecting airways of the lungs, which is characterized by recurring and variable narrowing of the small airways with resultant obstruction of the airflow that results in attacks of cough, wheeze, chest tightness and shortness of breath [3]. The characteristic airways hyper-responsiveness in asthma is initiated in response to provocative agents as irritant gases or pollens and symptoms are usually aggravated in the morning or at night [4].
Due to the fact that asthma is a chronic inflammatory disease, several studies related to bronchial asthma have focused on the mechanisms that control inflammation and are involved in the induction of immune tolerance to specific antigens [5].
Studies have postulated that the imbalance of T-helper cells (TH), especially Th1/Th2 imbalance might be the main factor responsible for the development of asthma [6]. Th1 cells are critical in defense mechanisms of cells and they play a major role in promoting proinflammatory immune reactions in response to infection as Th1 cells produce both IFN-γ and interleukin 2 (IL-2). On the other hand, Th2 cells mediate allergic inflammation and promote antibody-dependent immune responses as they generate several cytokines (IL-4, IL-5, IL-6, IL-9 and IL-13) [7].
Interleukin 18 (IL-18), which is one of the members of IL-1 family, was formerly known as IFN-γ inducing factor (IFGIF), and was also known to have an influence on the balance of Th1/Th2 immune response [8]. Evidence has demonstrated that IL-18 can induce IFN-γ production and hence promote Th1 responses, which is attained when IL-18 synergizes with IL-12 [9].
Nevertheless, IL-18 can promote Th2 responses through the production of IL-13. IL-18 can indirectly induce B-cell isotype switching to IgE production and can also induce the production of Th2 cytokines and hence it is involved in the development of allergic inflammatory reactions. Furthermore, IL-18 can synergize with IL-12 and inhibit IgE synthesis [10]. Moreover, Yoshimoto et al. reported that IL-18 may be critical for IgE production in vivo, providing a potential therapeutic target in allergic disorders [11].
IL-18 is located on chromosome 11q22.2-22.3, which is the region that encompasses several functional promoter region polymorphisms. In addition, large quantities of evidence support the hypothesis that IL-18 genetic polymorphism may cause abnormal expression of IL-18, which is responsible for mediating T-cell regulation leading to development of allergic diseases [12].
Allergic disorders have been linked to chromosome 11q22, which harbors the IL-18 gene, and IL-18 gene enhances IL-4/IL-13 production, which leads to the induction of IgE production [13]. Therefore, given the known importance of IL-18 (-607C/A) single nucleotide polymorphism (SNP) in the development of allergic diseases and the variation in the susceptibility to immune disorders in different ethnic groups, we set to investigate the possible genetic influence of IL-18-607C/A SNP on the development of asthma in children, its relation to disease severity and its effect on the serum level of IgE.
Material and methods
Patients
A total of 100 subjects of Egyptian children, including 60 patients (32 males and 28 females) were included in the study. The patients were recruited from the outpatient Allergy Clinic of Beni-Suef University Hospital, between January 2013 and February 2014. The ages of the patients ranged from 2 to 12 years. The patients were diagnosed by a pediatrician, and the diagnosis and classification of clinical severity were done according to the Global Initiative for Asthma (GINA) guidelines [14]. Patients with a history of chronic lung disease, non-atopic asthmatic children and patients treated with steroids within 2 weeks before sampling were excluded from the study. The patients group consisted of 20 children with mild persistent asthma, 30 children with moderate persistent asthma and 10 children with severe persistent asthma. The forty control subjects enrolled in the study consisted of 40 age and sex matched healthy children, who were free of any form of atopic and non-atopic manifestations and had no personal or family history of asthma or any other atopic manifestations. The guardians of the patients and control subjects were informed about the purpose of the study and consent was taken from all the participants’ guardians. The study was approved by the Research Ethical Committee of the Faculty of Medicine, Beni-Suef University (FMBSU REC) which operates according to the Guidelines of the Declaration of Helsinki, International Conference of Harmonization ICH, and United States Codes of Federal Regulations registered in the office of Human Protection under FWA#: FWA00013564.
Specimen collection
Approximately 6 ml of venous blood were withdrawn from all subjects under aseptic conditions. Two ml were collected in an EDTA vacutainer for complete blood picture (CBC) analysis. For determination of total serum IgE level, 2 ml of blood were collected in a plain tube with no anticoagulant and 2 ml of blood were collected in a sterile EDTA vacutainer for DNA extraction.
Detection of total serum IgE
Total serum IgE was measured using an Elisa Kit (Chemux Bioscience, Inc.), and the optical density was measured at 450 nm using and Elisa plate reader (Stat Fax). The total IgE concentration was expressed in IU/l according to the curves obtained from the standards.
Genotyping
Genomic DNA was extracted from EDTA anti-coagulated whole blood using QIAamp DNA Mini Kit (Cat.no. 51104, QIAGEN) according to the manufacturer’s protocol. The sequence flanking IL-18-607C/A SNP was amplified by PCR. Genotyping of the SNP was determined by restriction fragment length polymorphism assay (RFLP) [15]. The 171-bp region of the IL-18 gene, encompassing the -607C/A polymorphism site, was amplified via polymerase chain reaction (PCR) using the sense: 5’-GCCCTCTTACCTGAATTTTGGTAGCCCTC-3’ and antisense: 5’-AGATTTACTTTTCAGTGGAACAGGAGTCC-3’ primer pair. The total reaction volume was 25 µl: 5 µl DNA, 12.5 µl Dream Taq green PCR master mix (Fermentas), 1 µl of each primer and 5.5 µl of nuclease-free water. The PCR was performed in T-personal thermal cycler (Biometra) under the following cycling conditions: initially, denaturation for 5 min at 95°C, followed by 30 cycles of amplification (30 s at 95°C, 30 s at 60°C and 30 s at 72°C). The elongation step for 10 min at 72°C. The PCR products were separated on 2% agarose gel using 1x TEA buffer for 40 min at 100 volts, followed by ultraviolet visualization.
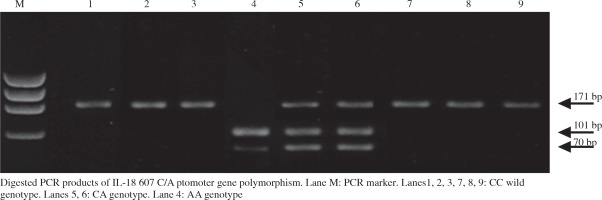
The PCR was followed by digestion with the restriction enzyme Tru1I (Thermo Scientific Cat. no.: #FD0984) according to the manufacturer’s protocol (Fig. 1). Digested PCR fragments were separated on 4% agarose gel electrophoresis stained with ethidium bromide, followed by ultra violet visualization. The PCR digested products generated a 171 bp un-cleaved fragment which represented the C allele, a 101 bp fragment and a 70 bp fragment which represented the A allele. To confirm the results of PCR-RFLP analysis, 10% of the samples were genotyped twice and the interpretation of the results was performed by two different observers and the results conformed to the original analysis.
Statistical analysis
The collected data were reviewed. Coding and statistical analysis was done by using SPSS program (Statistical Package for the Social Sciences; SPSS Inc., Chicago, IL, USA) version 16 for Microsoft Windows. Mean, median, range and standard deviation were calculated to measure central tendency and dispersion of quantitative data while frequency of occurrence was calculated to measure qualitative data. Student t test was used to determine the significance in difference between two means. χ2 test was done for comparison of qualitative data and Fisher’s exact test was used instead when the count of cell was less than 5. Odds ratios (ORs) with 95% confidence intervals (CI) were calculated whenever applicable, to test the association between the genotype and RA. Analysis of variance (ANOVA) test used to determine the difference between more than 2 means. The significance of the OR was calculated by a 2 by 2 contingency table. Kruskal-Wallis test was used for calculation of non-parametric data between groups. Genotype distributions were compared with those expected for samples from populations in Hardy-Weinberg equilibrium using a χ2 test (1 df). The level of significance was taken at p-value of < 0.05.
Results
In the current case control study, the study population comprised 60 children diagnosed with persistent asthma and 40 healthy control subjects. The percentage of female patients was 46.7%, while the percentage of male patients was 53.3%. The mean age of the patients was 6.5 ±3, while the mean age of the healthy control subjects was 7.3 ±3.1. In the current study, the serum level of IgE was compared in both asthmatic patients and healthy control groups. A statistically significant difference was detected in serum IgE levels between the 2 groups (p = 0.019). The demographic, clinical and laboratory data of the patients and the healthy control group are summarized in Table 1.
Table 1
Clinical and laboratory data of patients and control subjects
| Clinical and laboratory data | Asthmatic children (n = 60) | Healthy subjects (n = 40) | p-value |
|---|---|---|---|
| Age, mean ±SD | 6.5 ±3.0 | 7.3 ±3.1 | 0.382 |
| Sex, n(%) | 0.773 | ||
| Female | 32 (53.5) | 24 (46.7) | 24 (60) |
| Male | 24 (46.7) | 16 (40) | |
| Disease duration, years (mean ±SD) | 5.1 ±2.4 | – | |
| Severity of asthma, n(%) | |||
| Mild persistent | 20 (33.3) | ||
| Moderate persistent | 30 (50) | ||
| Severe persistent | 10 (16.7) | ||
| IgE, IU/ml (mean ±SD) | 135 ±59.5 | 48.4 ±21.1 | 0.019* |
| Total WBCs (mean ±SD) | 6.2 ±3.1 | 6.1± 4.3 | 0.924 |
| Absolute eosinophilic, count/mm3 (mean ±SD) | 280 ±33.2 | 8.8 ±5.3 | 0.001* |
The frequency distribution of IL-18-607C/A SNP genotypes in asthmatic children did not show deviation from Hardy-Weinberg Equilibrium (HWE) (p = 0.0859), similarly, the genotype frequency distribution of healthy control subjects conformed with HWE (p = 0.396).
Analysis of distribution of IL-18-607C/A SNP alleles showed that the allele frequency distribution for asthmatic children was 56.73% and 43.33% for C and A alleles, respectively. On the other hand, the frequency distribution for C and A alleles in healthy control subjects is 82.5% and 17.5%, respectively, and the difference between the two groups was statistically significant (p < 0.001) (Table 2).
Table 2
Genotype and allele frequencies of the IL-18-607C/A polymorphism in asthmatic children and healthy controls
| Variable | Asthmatic children (n = 60) | Healthy controls (n = 40) | OR (CI) | Significance test and p-value |
|---|---|---|---|---|
| Genotype, n (%) | χ2 = 18.3 p < 0.001* | |||
| CC | 16 (26.7) | 28 (70) | ||
| CA | 36 (60) | 10 (25) | ||
| AA | 8 (13.3) | 2 (5) | ||
| Genotype, n (%) | χ2 = 18.29 p < 0.001* | |||
| Dominant model (AA/AC) | 44 (73.3) | 12 (30) | 6.417 (2.432-17.289) | |
| Wild type (CC) | 16 (26.66) | 28 (70) | 1.00 | |
| Allele, n (%) | χ2 = 14.48 p < 0.001 | |||
| C | 68 (56.67) | 66 (82.5) | 1.00 | |
| A | 52 (43.33) | 14 (17.5) | 0.227 (0.132-0.574) |
Regarding the frequency distribution of the -607C/A (rs 1946518) genotypes in asthmatic children and control groups, the heterozygous CA genotype was significantly higher in asthmatic patients compared to the healthy control group (χ2 =18.3, p < 0.001). Moreover, in analysis of risk estimates between the SNP and the development of asthma in children we detected that asthmatic children carrying the AA/AC genotype of -607C/A SNP were associated with an increased risk of occurrence of asthma (OR = 6.417; CI: 2.432-17.289) (Table 2).
For the next analysis, in order to detect an association between the severity of asthma and rs 1946518, we stratified the asthmatic children into 3 groups according to the degree of severity of asthma in accordance with GINA guidelines (2011). However, we did not detect a statistically significant difference between rs 1946518 and the degree of severity of asthma in the patients’ groups (χ2 = 0.76, p = 0.94) (Table 3).
Table 3
IL-18-607C/A genotype frequency among asthmatic children according to the degree of clinical severity
Finally, analysis of the association between IL-18 (C/A, rs 1946518) genotypes and serum level of IgE in both patients and control groups revealed that the serum level of IgE was higher in asthmatic patients carrying the mutant heterozygous AA genotype compared to other asthmatic patients carrying the CA and CC genotypes respectively, however, the difference between groups did not reach statistical significance. On the other hand, the serum IgE level was the lowest in healthy control subjects carrying the CA genotype compared to controls carrying the CA and AA genotypes and the difference between the two groups was statistically significant (p = 0.046) (Table 4).
Table 4
Association between IL-18-607 (C/A) and serum levels of IgE in asthmatic children and in the healthy control group
| Variable | Median range of serum IgE (IU/l) in asthmatic patients | p-value | Median range of serum IgE (IU/l) in healthy controls | p-value |
|---|---|---|---|---|
| Genotype | ||||
| CC (n = 16) | 83.5 (40-101) | 0.054 | 59.6 (26.5-101) | 0.046* |
| CA (n = 36) | 82 (40.6-659) | 16 (4-73.9) | ||
| AA (n = 8) | 103 (101-457) | 41.95 (38.4-45.5) |
Discussion
Asthma has been related to blood or airway eosinophilia. Asthmatic patients with high eosinophilic counts were found to be at risk for more severe disease. Eosinophils may actively lead to innate and adaptive immune responses with resultant inflammatory cascades through the release of different cytokines, growth factors and lipid mediators [16].
In the current study, the absolute eosinophilic count was significantly higher in the asthmatic children compared to the healthy control group (p < 0.001). This conformed with the results of El-Zohery et al. who tested eosinophilic count as one of the five biochemical markers in asthmatic children’s blood as parameters for bronchial asthma induction due to house-dust mite allergy [17]. This could be explained by the fact that in cases of atopic asthma, Th2 cells are activated with a subsequent release of IL-15 with the resultant enhancement of eosinophilia [18].
The causal relationship detected between asthma and allergens depends on epidemiologic findings, demonstrating a strong association between total IgE and asthma [19].
In this regard, we investigated the serum level of IgE and we detected that the total serum IgE was significantly higher in asthmatic children compared to the healthy subjects (p = 0.019). Our results were in consistency with the results of the study conducted by Obaidi et al. which reported that the serum IgE level was higher in asthmatic patients compared to the control group and they postulated that the serum IgE level was predictive of asthma and might be used to differentiate between asthmatic and non-asthmatic individuals in association with other biomarkers [19]. Our results were also in accordance with the study conducted by Satwany et al., which postulated that the total IgE level is a strong indicator of allergy in asthmatic children [20].
Currently, IL-18 genetic polymorphisms have been postulated to be involved in the development of allergic diseases as it has been postulated that IL-18 genetic polymorphisms may cause abnormal expression of IL-18, which is responsible for mediating the T-cell regulation, thereby leading to the pathological development of allergic diseases. The results of these studies have been emphasized by genetic investigations, revealing the association of polymorphisms within the gene with the occurrence of these diseases [12, 21].
The results of the current study were in accordance with the study conducted by Lachheb et al., which revealed a significant association in allele and genotype frequencies in both asthmatic and non-asthmatic children [8]. Our results were in consistency with the results of the meta-analysis conducted by Ma et al. which revealed that IL-18-607C/A polymorphism was associated with an increased risk of asthma in the recessive model, thus postulating that the C allele of IL-18C/A SNP might have a role in the development of asthma [7].
The above results could be explained by the fact that the allele variants of cytokine genes which are associated with promoter region SNPs, do not affect the protein amino-acid sequence but can impact the production of the cytokine, which might result in alteration of the immune response produced by the cytokine [12]. IL-18-607C/A promoter polymorphism is associated with the low transcriptional activity. The low promoter activity is associated with the -607A allele while the higher promoter activity was associated with the C allele. The low promoter activity could be affected by a change from C to A position at -607 which resulted in disruption of a potential cAMP-responsive element binding protein binding site, which leads to a decrease in production of IL-18 [8, 22].
However, the results of the present study were not in consistency with the results of the meta-analysis conducted by Cheng et al., which did not find an association between IL-18-607C/A SNP and the risk of allergic diseases in the overall analysis [12]. Further on, our results were not in agreement with the results of the studies carried out by Heinzmann et al. which did not detect an association between asthma and the SNP [21].
In the current study, we stratified the asthmatic patients into 3 groups according to disease severity and we analyzed the presence of an association between the degree of disease severity and IL-18-607C/A SNP. Although the minor A allele was higher in mild and moderate disease severity groups, we could not detect a statistically significant difference between the degree of asthma severity and the SNP. Our results were in accordance with the study conducted by Izakovicova Holla et al., which was conducted in adult asthmatic Czech patients and did not detect an association between IL-18-607C/A polymorphism and the degree of severity of asthma [23]. Further on, Lachheb et al. was unable to detect a relation between IL-18-607 SNP genotypes and alleles with the severity of asthma [8]. However, Pawlik et al. indicated that patients carrying the CC genotype were significantly more frequent among patients with severe asthma [24]. However, our results regarding the association of the SNP with disease severity should be considered as a preliminary report for further studies due to the small sample number used in the analysis.
Cardon and Bell indicated that some cytokine SNPs are associated with allergies, asthma or elevated IgE levels [25]. However, our results regarding the association of IL-18-607C/A SNP and the serum level of IgE, were in agreement with the results of the study conducted by Lacheb et al., which did not detect a significant association between genotype frequency distribution and IgE level in asthmatic patients [8]. On the other hand, our results were in disagreement with the results of the study conducted in Germany on patients with hay fever and atopy, which postulated an association between IL-18 polymorphisms and the level of IgE [26].
The discordance between some of the results of the current study and the results of studies conducted in different populations might be attributed to a variety of factors. Generally, an allergic disease is a multifactorial disease controlled by interplay of several susceptibility genes and environmental factors and hence, no single gene or environmental factor can have a large effect on the susceptibility to allergic diseases [22]. Further on, conflicting results might be caused by the differences in geographical regions with differences of ethnic backgrounds of populations in which the studies were conducted. Moreover, the differences in genotyping methods used for detection of the SNP and the differences in sample sizes enrolled in different studies might be the reason for inconsistent results.
Conclusions
The current study postulates that both allele and genotype frequency distribution of IL-18-607C/A SNP are significantly associated with asthma in our study group. There is no statistically significant association between the degree of severity of asthma and -607C/A genotype and allele frequency distribution despite the higher frequency of the minor A allele in mild and moderate disease severity groups. Moreover, the total serum levels of IgE in asthmastic children carrying the CA genotype was significantly higher than the control group. However, due to the small sample size included in the current study and clinical heterogeneity, which have resulted in lowering the power of the study, the results of the current study should be treated cautiously and further studies with larger population samples to establish the genetic association are needed.